AI for Biology: What's Actually Working in 2026

A few years ago, “AI in biology” was mostly a pitch deck topic. Foundation models were reshaping natural language processing and image generation, but biology remained largely untouched. The data was too messy, the domains too specialized, the validation cycles too slow.
That’s changed. Not everywhere, and not all at once, but in specific areas the results are now tangible: drugs designed by AI entering late-stage clinical trials, protein design tools producing experimentally validated structures, genomic models processing millions of base pairs with single-nucleotide resolution, and diagnostic systems outperforming experienced clinicians on rare disease cases.
This post maps out where AI is delivering real results in biology today, where the progress is genuine but early, and where the gap between promise and practice remains wide.
AI in Genomics: From Sequencing to Understanding
Sequencing a genome is no longer the hard part. The cost of whole-genome sequencing has dropped below $200, and GPU-accelerated pipelines have compressed germline variant calling from 16 hours to under five minutes. The challenge has shifted from generating genomic data to interpreting it.
Foundation models for the genome
Two large-scale genomic foundation models have emerged that treat DNA itself as a language to be modeled.
AlphaGenome, published by Google DeepMind in Nature in January 2026, takes up to one million base pairs of DNA as input and predicts thousands of molecular properties at single-base-pair resolution: gene expression levels, chromatin accessibility, histone modifications, transcription factor binding, and splice site locations. In head-to-head evaluations, it matched or exceeded existing specialized models in 25 of 26 variant effect prediction tasks. Nearly 3,000 scientists across 160 countries have started using the open-source tool.
Evo 2, developed by the Arc Institute, UC Berkeley, and NVIDIA, is a 40-billion-parameter model trained on over 9.3 trillion nucleotides from 128,000+ whole genomes spanning all domains of life. It processes sequences up to one million nucleotides long with single-nucleotide resolution, and it has been released as open-source software.
These models don’t replace clinical variant interpretation, but they provide a new layer of functional prediction that can inform whether a given variant is likely to disrupt gene regulation, protein expression, or splicing.
Clinical-scale variant interpretation
On the applied side, AI is starting to make variant interpretation faster and more consistent at clinical scale. GeneDx, one of the largest clinical genomics laboratories, now works with a dataset of nearly one million exomes and genomes, roughly one million uniquely classified variants, and over seven million phenotypic data points. Their AI-based variant ranker has reduced the manual interpretation workload while maintaining recall of pathogenic variants, and their diagnostic yield stands at 51.4% across more than 300,000 cases.
The broader challenge remains: most variants identified in clinical sequencing are still classified as variants of uncertain significance (VUS), and resolving them requires synthesizing evidence across multiple databases, structural models, and clinical context. This is the problem that molecular intelligence platforms are built to address, combining database access, structural reasoning, and AI interpretation in a single workspace rather than requiring clinicians to manually cross-reference disconnected tools.
AI in Protein Structure and Design
The 2024 Nobel Prize in Chemistry went to Demis Hassabis and John Jumper for AlphaFold, and to David Baker for computational protein design. It was a recognition of work that has already reshaped how biology approaches structural questions.
Beyond static structure prediction
AlphaFold solved the protein structure prediction problem for single chains with remarkable accuracy. AlphaFold 3, published in Nature in May 2024, extended this to protein complexes involving DNA, RNA, small-molecule ligands, and ions, with at least a 50% improvement in accuracy for protein-ligand interactions compared to previous methods. As of late 2025, the AlphaFold 3 paper had been cited over 9,000 times.
But predicting a structure is only the beginning. The field has moved from asking “what does this protein look like?” to “can we design a protein that does something specific?”
Generative protein design
In December 2025, David Baker’s lab released RFdiffusion3, described as their most versatile protein engineering tool to date. Unlike earlier tools that worked at the residue level, RFdiffusion3 designs proteins at the all-atom level, enabling it to create proteins that interact with virtually every type of molecule in cells: DNA, other proteins, small molecules, and even catalytic active sites.
The tool has been experimentally validated. Designed proteins bind target ligands with expected geometry, recognize specific DNA shapes, and show catalytic activity. It consolidates capabilities that previously required multiple specialized tools into a single open-source platform.
For teams working in drug discovery or protein engineering, this means the feedback loop between design and validation is getting shorter. A researcher can now generate candidate protein binders computationally, predict their structures, assess their stability, and prioritize the most promising designs for experimental testing. Platforms that integrate protein structure prediction with broader biological analysis make this workflow more accessible to teams without deep computational expertise.
AI in Drug Discovery: Clinical Evidence Is Arriving
AI-designed drugs are no longer theoretical. They’re in clinical trials, and some are producing results.
The pipeline is real
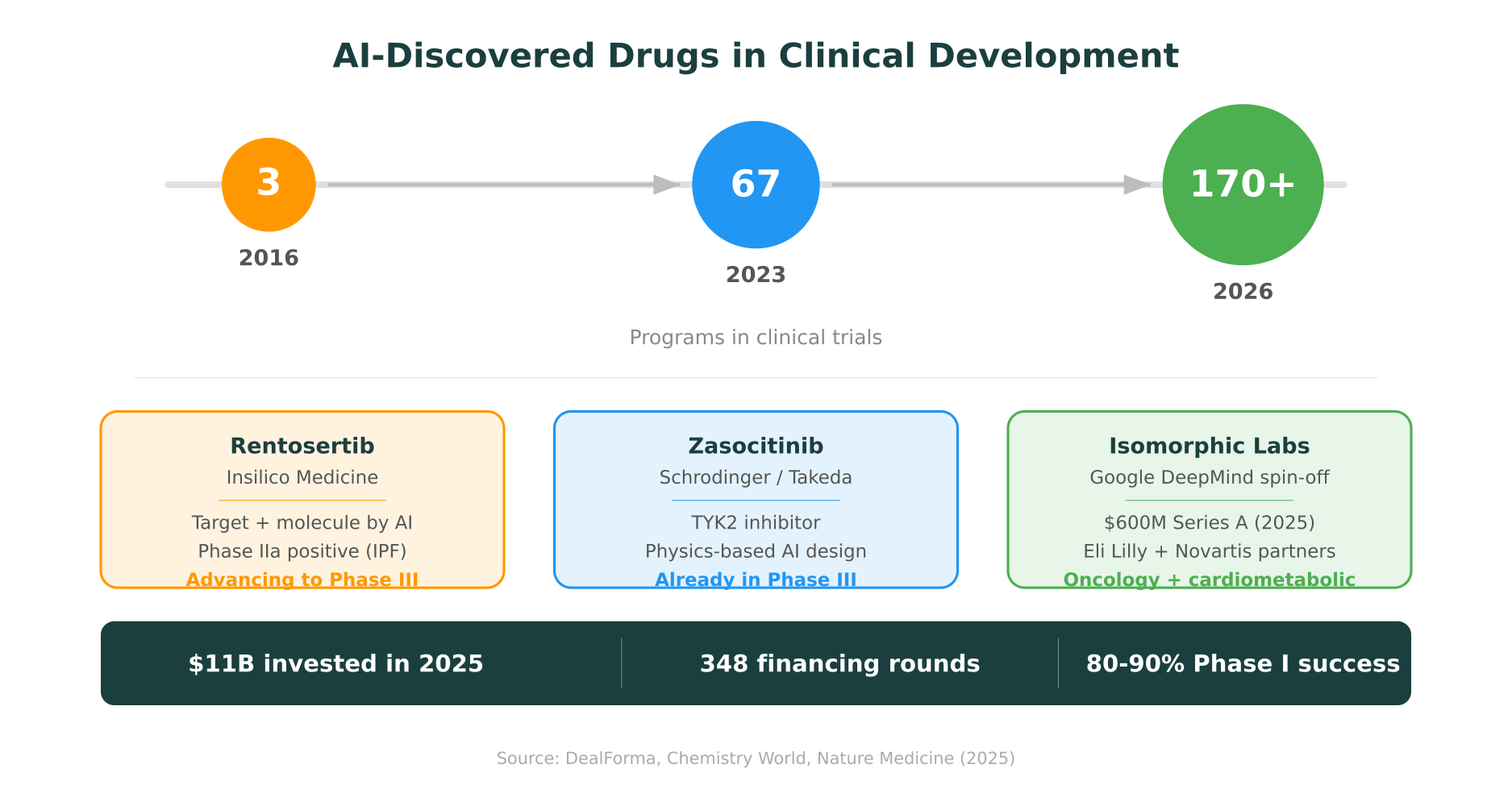
As of early 2026, over 170 AI-discovered drug programs are in clinical development, up from roughly 67 in 2023 and just 3 in 2016. Early data has been encouraging: AI-discovered molecules have shown Phase I success rates in the range of 80-90%, which is notably above historical industry averages, though the sample sizes are still small and selection bias is a factor.
No AI-designed drug has received FDA approval yet. That milestone is likely still a year or more away. But several programs have advanced far enough to be genuinely informative.
Notable programs
Insilico Medicine’s rentosertib is the most closely watched. Both the drug target (TNIK) and the molecule itself were identified and designed using generative AI. Phase IIa results for idiopathic pulmonary fibrosis, published in Nature Medicine in 2025, were positive. The program is advancing toward Phase III, which would make it the first fully AI-designed drug to reach that stage. Sanofi signed a collaboration with Insilico worth up to $1.2 billion.
Zasocitinib, a TYK2 inhibitor designed using Schrodinger’s physics-based AI platform, is already in Phase III trials (developed through Nimbus Therapeutics and now Takeda).
Isomorphic Labs, a Google DeepMind spin-off, raised a $600 million Series A in March 2025 and has partnerships with Eli Lilly and Novartis. They’re advancing their own oncology and cardiometabolic programs toward first-in-human studies.
The investment landscape reflects the momentum: 348 financing rounds raised $11 billion in AI-driven drug discovery in 2025 alone. Chai Discovery, backed by OpenAI, raised $130 million at a $1.3 billion valuation. Venture capital into generative protein design platforms tripled compared to 2024.
What this means practically
The shift isn’t that AI replaces medicinal chemists or clinical researchers. It compresses the early stages of discovery. Target identification, hit generation, and lead optimization, steps that traditionally took years of iterative design-make-test-analyze cycles, can now be done in months. The clinical trial process itself hasn’t changed, but the molecules entering it are arriving faster and, in some cases, with better predicted properties.
This connects directly to how molecular intelligence accelerates the drug discovery pipeline: by combining structural prediction, molecular property modeling, and database reasoning, AI platforms can evaluate more candidates computationally before a single molecule is synthesized.
AI in Disease Diagnosis: Rare Diseases as a Proving Ground
Rare diseases affect roughly 300 million people worldwide, and the average time to diagnosis is over five years. Many patients see multiple specialists before receiving a correct diagnosis, and some never do. This is an area where AI’s ability to process large amounts of heterogeneous clinical data is starting to make a measurable difference.
DeepRare
In February 2026, a system called DeepRare was published in Nature. It uses an agentic AI architecture, combining a central LLM with over 40 specialized tools for phenotype extraction, disease normalization, knowledge retrieval, and genotype analysis.
Evaluated on 6,401 cases from multiple global medical centers, DeepRare achieved a 64.4% first-attempt diagnostic accuracy, compared to 54.6% among physicians with over 10 years of experience. With multi-modal inputs (combining phenotype and genotype data), recall reached 70.6%.
The system has been deployed on an online platform since July 2025, with over 600 medical institutions registered worldwide.
Why rare diseases are a natural fit
Rare disease diagnosis is essentially a pattern-matching problem across an enormous feature space. A single patient may present with a constellation of symptoms that maps to any of thousands of possible genetic conditions. No individual clinician can hold the full landscape of 7,000+ rare diseases in working memory, but an AI system can systematically compare a patient’s phenotype against all known disease signatures.
The limitation is that diagnostic accuracy, even at 64%, still means roughly one in three cases is missed on first attempt. These systems work best as decision support, narrowing the differential and surfacing diagnoses that a clinician might not have considered, rather than as standalone diagnostic tools.
AI in Multi-Omics: Connecting the Layers
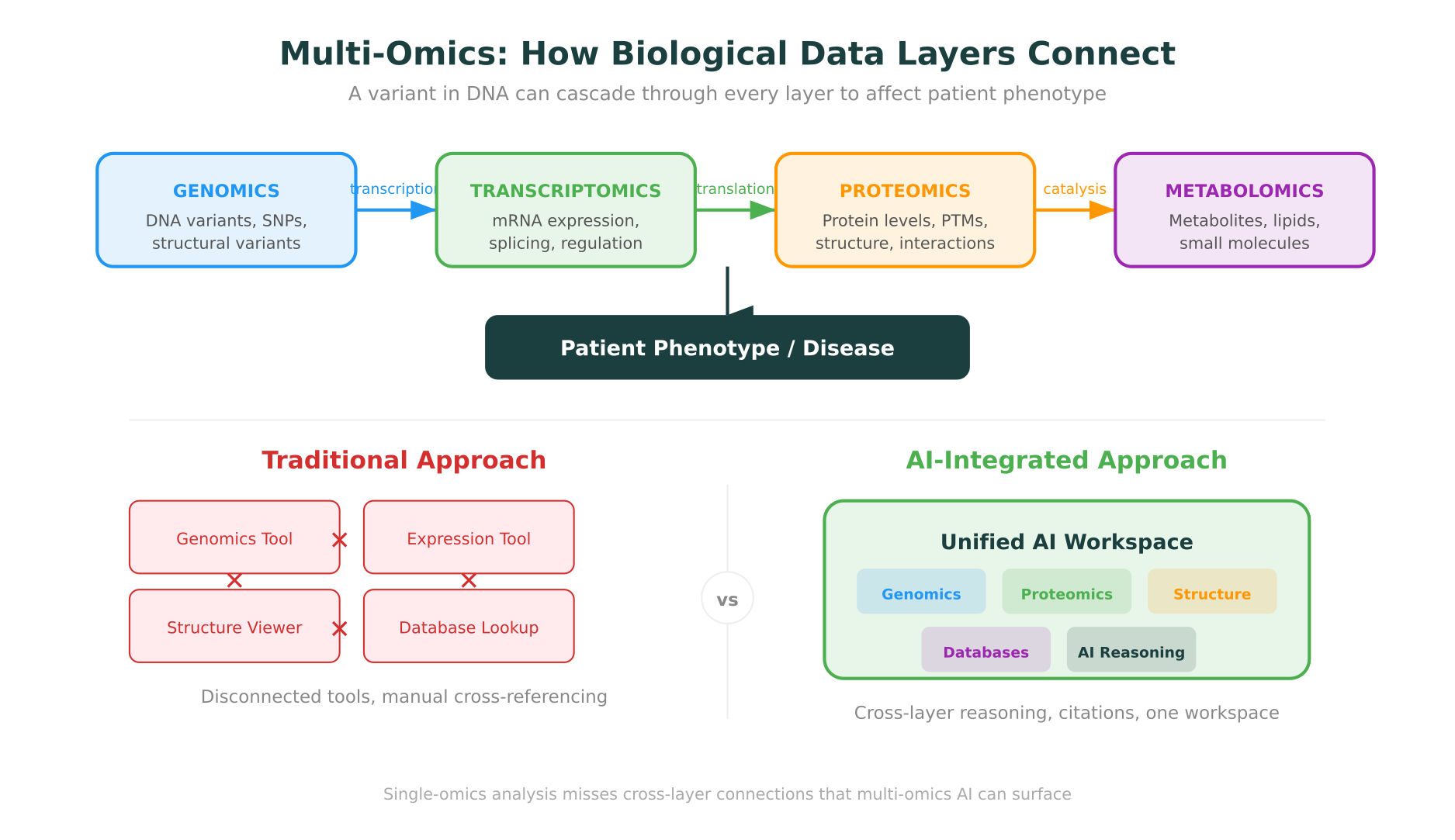
Most biological questions don’t live within a single data type. A genomic variant might alter gene expression, which might change protein levels, which might disrupt a metabolic pathway. Understanding this cascade requires integrating data across genomics, transcriptomics, proteomics, and metabolomics, what the field calls multi-omics analysis.
AI is making this integration more tractable, though the field is still early.
Where things stand
Recent work has focused on building end-to-end deep learning models that explicitly model the flow of information from DNA to RNA to protein to metabolite to phenotype. Molecule foundation models generate embeddings for molecular perturbations, while cell foundation models extract features from each omics layer using unlabeled data.
AI-powered mass spectrometry is also transforming the proteomics and metabolomics side of the equation, improving the speed and accuracy of protein identification and metabolite annotation.
The practical challenge is data. Multi-omics experiments are expensive, and paired datasets (where the same samples have been profiled across multiple omics layers) are still relatively scarce. Labeled training data is even scarcer. This limits how quickly supervised AI models can improve.
What multi-omics AI enables
When it works, multi-omics AI can surface connections that single-omics analysis misses. A variant that looks benign based on population frequency alone might become significant when correlated with abnormal expression levels and disrupted protein interactions. A drug target that looks promising from genomic data alone might reveal off-target effects visible only in the metabolomic layer.
This is an area where having a unified workspace matters. Switching between separate tools for genomic annotation, expression analysis, and protein structure visualization introduces friction and increases the chance of missing cross-layer connections. Platforms that bring multi-omics analysis into a single environment, with integrated databases and AI reasoning, are better positioned to surface these insights.
What Ties It All Together
The common thread across genomics, protein design, drug discovery, diagnostics, and multi-omics isn’t any single algorithm or model. It’s a shift in how biological research gets done.
Biology is becoming a field where the limiting factor is no longer data generation or even computational power. It’s the ability to reason across heterogeneous data types, structured databases, and domain-specific knowledge in a coherent way. The tools that are producing real results in 2026 share a few characteristics:
They’re grounded in domain-specific data. AlphaGenome works because it was trained on genomic data, not internet text. RFdiffusion3 works because it models all-atom physics, not just residue-level abstractions. DeepRare works because it connects to 40+ specialized biological tools, not because it has a larger language model.
They integrate rather than isolate. The most impactful tools connect data types that were previously analyzed in separate workflows. Variant interpretation that incorporates protein structure. Drug discovery that combines target prediction with molecular design. Diagnostics that merge phenotype data with genotype analysis.
They show their reasoning. In biology, an answer without evidence is just a guess. The tools gaining clinical and research adoption are the ones that provide citations, surface uncertainties, and make their reasoning auditable. This is especially critical in clinical genomics and drug development, where decisions have direct consequences for patients.
This is the design philosophy behind molecular intelligence: not a single AI model, but an intelligent infrastructure that connects databases, structural tools, code execution, and domain reasoning into one workspace where biology teams can move from question to evidence-backed answer without stitching together disconnected tools.
The field is still early. Most of these tools are measured in months of availability, not years. Validation datasets are growing but remain limited in some areas. And the gap between what AI can do in a controlled benchmark and what it reliably does in clinical or research practice is real.
But the trajectory is clear. AI for biology is no longer a future promise. It’s a present reality, uneven and incomplete, but producing results that matter.
MIP is Purna AI’s Molecular Intelligence Platform, an AI-powered workspace for biology teams. Variant interpretation, protein structure prediction, code execution in containerized environments, and 30+ database integrations in one place. Explore the platform at purna.ai.
Explore Purna's Molecular Intelligence Platform
AI-powered workspace for biology teams to accelerate drug discovery from target identification to lead optimization.
Try Purna AI →
