How to Read a Genomic Report: A Guide for Clinicians

Genomic testing is no longer confined to genetics clinics. Oncologists order tumor panels. Cardiologists request inherited arrhythmia screens. Neurologists evaluate pharmacogenomic profiles before prescribing. Pediatricians receive whole exome results for children with developmental delays. In all of these cases, the clinician who ordered the test receives a genomic report, and that report needs to be interpreted correctly for the patient’s care to benefit.
The challenge is that most clinicians who order or receive genomic tests are not genetics specialists. A 2021 survey published in Genetics in Medicine found that fewer than 30% of non-geneticist physicians felt confident interpreting genomic reports independently. That gap between test ordering and report understanding creates real clinical risk: actionable findings may be overlooked, or, just as dangerously, uncertain findings may be treated as definitive diagnoses.
This guide explains how to read a genomic report section by section. It covers what each part means, how variant classifications work, when results should change clinical management, and when they should not. It is written for internists, oncologists, cardiologists, pediatricians, and any clinician who encounters genomic reports in practice.
Why Every Clinician Needs to Understand Genomic Reports
The volume of genomic testing in clinical medicine has grown substantially over the past decade. According to data from the National Institutes of Health, the number of available genetic tests exceeded 75,000 by 2024, spanning oncology, cardiology, neurology, reproductive medicine, and pharmacogenomics. This growth means that clinicians across specialties now encounter genomic data routinely, not just in rare disease or cancer genetics contexts.
Understanding how to read a genomic report matters for several reasons:
- Clinical action depends on correct interpretation. A pathogenic variant in BRCA1 on an oncology panel has direct implications for surgical planning, screening intervals, and family risk assessment. Missing this finding, or misinterpreting a VUS as pathogenic, can both lead to harm.
- Patients ask questions. When a patient receives genetic test results, they often turn to their primary care physician or specialist for explanation, not the testing laboratory.
- Referral decisions require baseline understanding. Knowing when to refer to a genetic counselor or medical geneticist requires recognizing which results are straightforward and which are complex.
- Reports vary across laboratories. Different labs present results in different formats, use different levels of detail, and may classify the same variant differently. A clinician who understands the underlying structure can navigate these differences.
Anatomy of a Genomic Report
Every genomic report, regardless of the testing laboratory, follows a general structure. The specific layout varies, but the core sections are consistent. Understanding this anatomy is the first step in reading any report confidently.
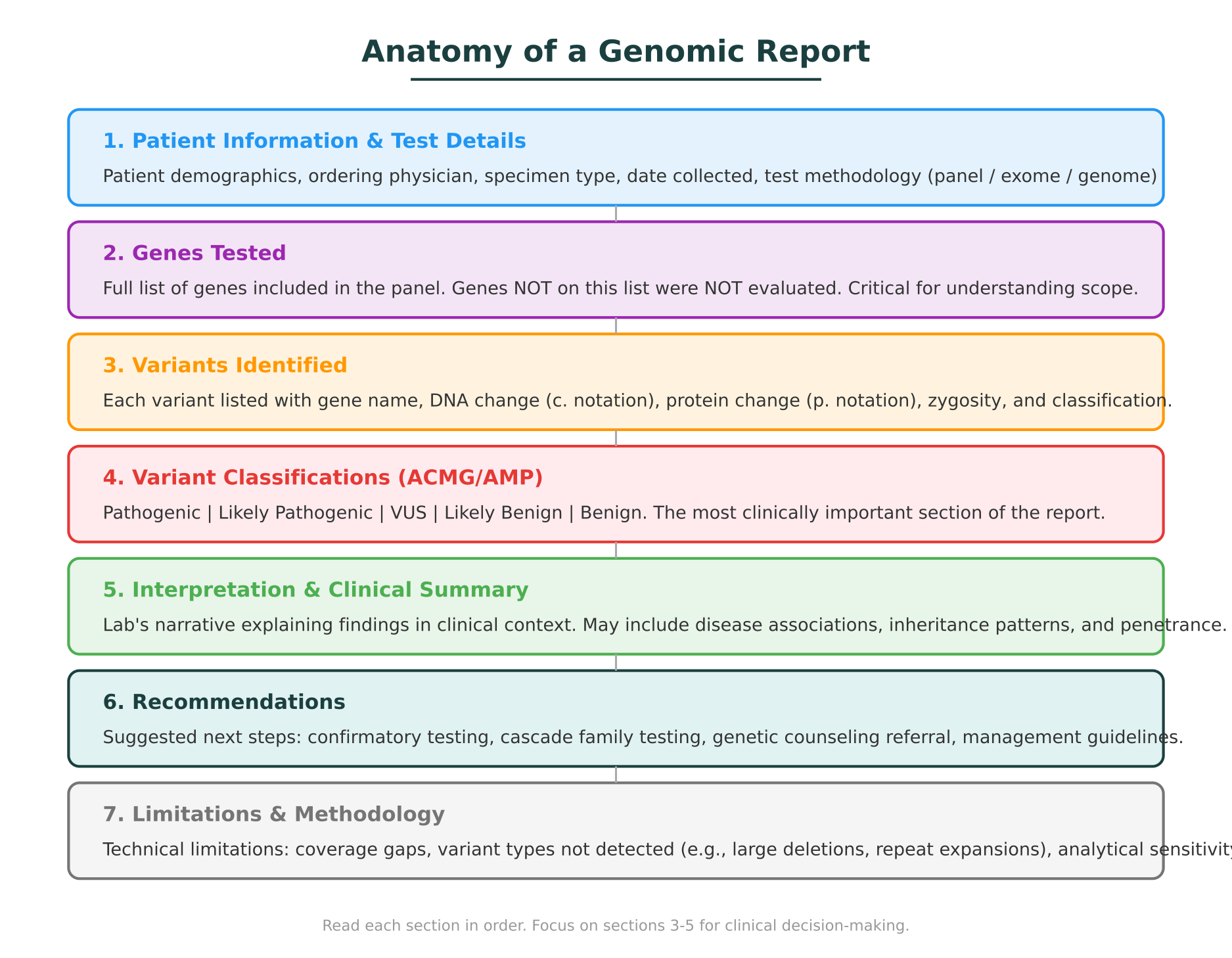

Patient information and test details
The top of the report identifies the patient, the ordering clinician, the specimen type (blood, saliva, tumor tissue), the date collected, and the date reported. It also specifies the test performed. This matters because the type of test determines the scope of what was evaluated:
- Gene panel: Tests a defined set of genes (anywhere from 5 to 500+) associated with specific conditions. Genes not on the panel were not evaluated.
- Whole exome sequencing (WES): Sequences the protein-coding regions of all ~20,000 genes. Broader than a panel but does not cover non-coding regions.
- Whole genome sequencing (WGS): Sequences the entire genome, including non-coding regions. The most comprehensive test, but interpretation of non-coding variants remains limited.
Genes tested
This section lists every gene included in the analysis. It is one of the most important sections for understanding what the test can and cannot tell you. If a gene is not on this list, no conclusion can be drawn about that gene from this report. A negative result means no reportable variants were found in the genes tested, not that the patient has no genetic risk.
Variants identified
This is the core findings section. Each variant is listed with standardized notation:
- Gene name (e.g., BRCA2, KCNQ1, TP53)
- DNA change in HGVS nomenclature (e.g., c.5946del)
- Protein change (e.g., p.Ser1982ArgfsTer22)
- Zygosity: heterozygous (one copy) or homozygous (two copies)
- Classification: the ACMG/AMP tier assigned to this variant
Some reports also include the genomic coordinates, the exon number, and the inheritance pattern of the associated condition (autosomal dominant, autosomal recessive, X-linked).
Interpretation and clinical summary
The laboratory provides a narrative section explaining the clinical significance of each finding. This includes the disease association of the gene, the evidence supporting the variant’s classification, and any relevant literature references. For pathogenic or likely pathogenic variants, this section typically includes information about penetrance, recommended management guidelines, and implications for family members.
Recommendations
Many reports include explicit next-step recommendations. These might include confirmatory testing by a second method (such as Sanger sequencing), cascade genetic testing for at-risk family members, referral to genetic counseling, or condition-specific surveillance protocols.
Limitations
Every genomic test has limitations. This section discloses what the test cannot detect. Common limitations include:
- Structural variants: Large deletions, duplications, and inversions may not be detected by sequencing-based methods.
- Repeat expansions: Conditions like Huntington disease or fragile X syndrome require specialized assays.
- Deep intronic variants: Variants outside the coding regions and splice sites are typically not captured by panels or exome sequencing.
- Mosaicism: Low-level mosaicism may fall below the detection threshold.
Reading this section helps prevent false reassurance from a “negative” result.
Understanding Variant Classifications: The ACMG/AMP System
The classification of each variant is the single most important piece of information on a genomic report for clinical decision-making. Nearly all clinical laboratories in the United States and many internationally use the framework published by the American College of Medical Genetics and Genomics (ACMG) and the Association for Molecular Pathology (AMP) in 2015 (Genetics in Medicine, Richards et al.).
The system assigns each variant to one of five tiers:
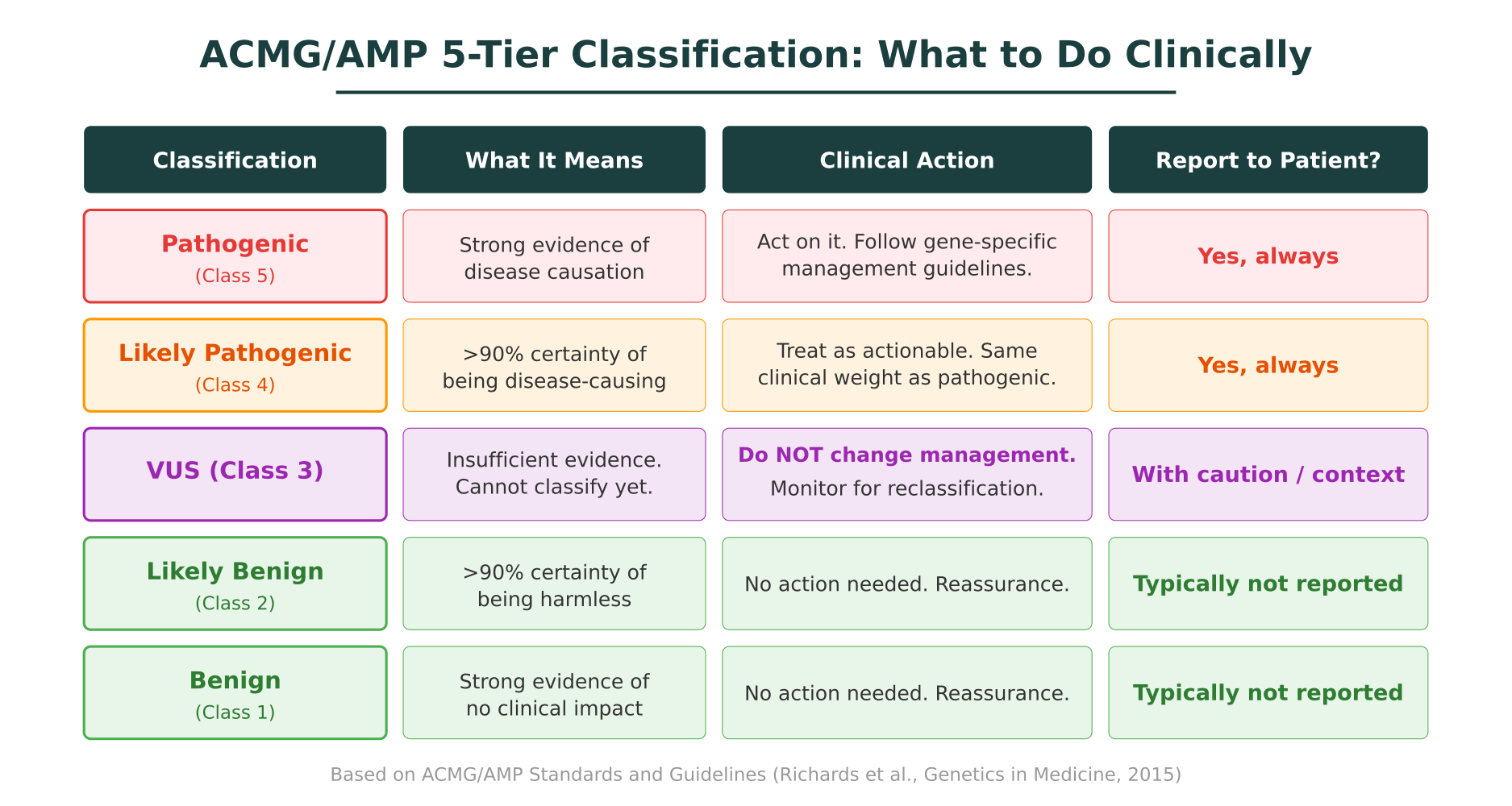

Pathogenic (Class 5)
Strong evidence supports that this variant causes disease. Evidence may include well-established functional studies, multiple affected families, absence in healthy populations, and computational predictions that all converge on a damaging effect. A pathogenic variant in a clinically relevant gene is an actionable finding.
Likely pathogenic (Class 4)
The evidence supports disease causation with greater than 90% certainty, but falls slightly short of the threshold for “pathogenic.” From a clinical management standpoint, likely pathogenic variants are treated with the same weight as pathogenic variants. ACMG guidelines explicitly state that both pathogenic and likely pathogenic variants should be used for clinical decision-making.
Variant of uncertain significance (Class 3)
A variant of uncertain significance (VUS) is the classification that causes the most confusion in clinical practice. A VUS means the laboratory found a variant, but the available evidence is insufficient to determine whether it is disease-causing or benign. This is not a diagnosis. A VUS should not, on its own, change clinical management.
VUS results are common. In ClinVar, approximately 50% of unique variant submissions carry the VUS label. The proportion is even higher for patients from underrepresented ancestral backgrounds, where population reference data is sparser.
Likely benign (Class 2) and Benign (Class 1)
These variants have strong evidence supporting that they do not cause disease. Most laboratories do not report likely benign and benign variants on clinical reports, as they are not clinically actionable. If they do appear, they require no action.
When to Act on Genomic Report Results
The classification tier directly determines whether and how to act. Understanding this mapping is essential for clinicians who receive genomic reports.
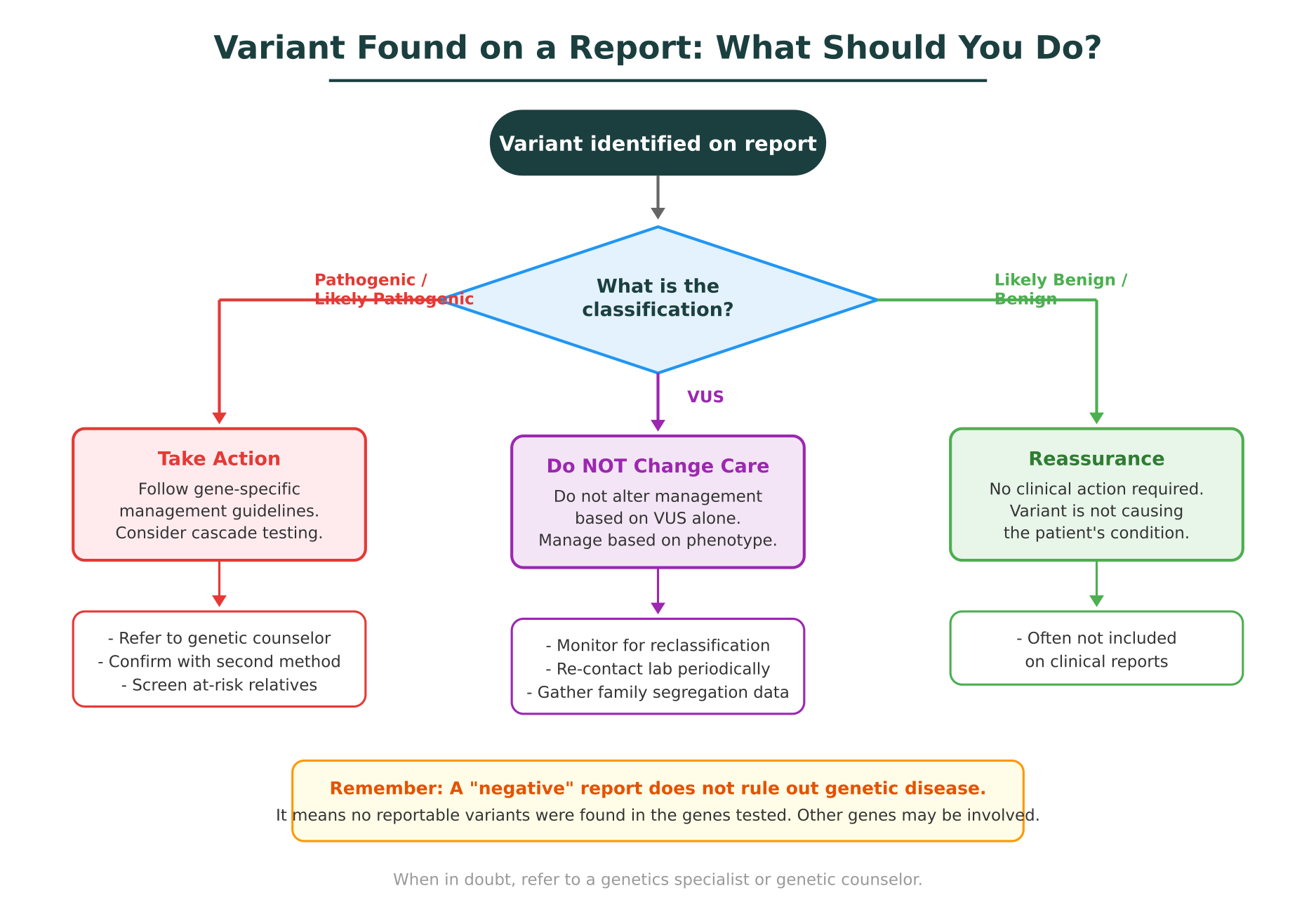
Pathogenic or likely pathogenic variants
When a report identifies a pathogenic or likely pathogenic variant in a gene relevant to the patient’s clinical context, this is an actionable finding. The appropriate response depends on the specific gene and condition:
- Follow published management guidelines. For many genes, professional societies have published evidence-based management recommendations. For example, NCCN guidelines outline screening and risk-reduction strategies for BRCA1/2 carriers. The Heart Rhythm Society provides management protocols for inherited arrhythmia genes like SCN5A and KCNQ1.
- Consider confirmatory testing. Some laboratories recommend confirming the variant by an orthogonal method, particularly for unexpected findings or variants detected at low allele fractions in tumor samples.
- Offer cascade testing to at-risk relatives. Once a causative variant is identified in a proband, targeted testing of family members is efficient, inexpensive, and clinically valuable. For a deeper look at how genomic findings translate into concrete clinical management steps, see From Variant to Visit: How Genomics Actually Changes Management.
- Refer to genetic counseling. Even when the result seems straightforward, genetic counseling helps patients understand inheritance patterns, reproductive implications, and psychological impact.
VUS: do not change management
This is the most critical point for non-genetics clinicians. A VUS should not be used as the basis for clinical decisions such as prophylactic surgery, changes in screening intervals, medication adjustments, or cascade family testing. The evidence is not there yet.
What to do with a VUS:
- Continue managing the patient based on their phenotype, family history, and clinical presentation, not the VUS.
- Note the VUS in the medical record so it can be revisited if the classification changes.
- Consider periodic recontact with the testing laboratory (every 1 to 2 years) to check whether the variant has been reclassified. Reclassification rates vary, but a 2019 analysis in Genetics in Medicine found that approximately 10% of VUS are reclassified within five years, most often to likely benign or benign.
- Refer to genetics if the clinical suspicion remains high despite the VUS. A genetic counselor or medical geneticist can evaluate whether additional evidence (segregation analysis, functional studies, updated database queries) might clarify the variant’s significance.
Negative result
A negative report means no pathogenic, likely pathogenic, or VUS results were identified in the genes tested. It does not mean the patient has no genetic risk. There may be causative variants in genes not included on the panel, in non-coding regions, or in variant types the test cannot detect (structural variants, repeat expansions). If clinical suspicion for a genetic condition remains strong after a negative result, consider broader testing (exome or genome sequencing) or referral to genetics.
Common Misunderstandings When Reading Genomic Reports
Several misunderstandings recur among clinicians who are not genetics specialists. Recognizing these can prevent errors in patient care.
”Negative” means no genetic risk
As discussed above, a negative result reflects only the scope of the test. Genes not tested, variant types not detectable, and regions not covered all represent blind spots. A negative result on a 50-gene panel does not rule out a genetic cause if the causative gene was not on the panel.
VUS means “probably pathogenic”
The word “uncertain” can be misleading. Some clinicians interpret a VUS as a likely positive finding, especially when it appears in a gene associated with their patient’s condition. In reality, data from large reclassification studies shows that most VUS that are eventually reclassified move toward benign, not pathogenic. Treating a VUS as pathogenic leads to unnecessary interventions, unwarranted anxiety, and inappropriate cascade testing.
All pathogenic variants require immediate intervention
A pathogenic variant is a confirmed genetic finding, but the clinical response depends on the specific gene, the condition’s penetrance, and the patient’s clinical context. Some pathogenic variants are associated with conditions that have reduced penetrance, meaning not all carriers develop disease. Others may be relevant to pharmacogenomics (affecting drug metabolism) rather than disease risk. The presence of a pathogenic variant does not always mean the patient will develop the associated condition.
The report is the final word
Variant classification is not static. As new evidence accumulates, including population data, functional studies, and case reports, variants can be reclassified. A variant classified as VUS today may be downgraded to likely benign next year, or upgraded to likely pathogenic. This is why noting VUS results in the medical record and periodically checking for updates matters.
Two labs always agree
Different laboratories may classify the same variant differently. This happens because laboratories may weigh evidence criteria differently, have access to different internal datasets, or interpret the same published data differently. A 2022 analysis in Genetics in Medicine documented discordance rates of approximately 10 to 15% for variant classifications across laboratories. When a classification seems inconsistent with the clinical picture, a second opinion from another laboratory or a genetics specialist may be warranted.
When to Refer to a Genetics Specialist
Not every genomic report requires genetics referral, but several situations warrant it:
- Multiple VUS in clinically relevant genes. Interpreting multiple uncertain findings in the context of a patient’s phenotype requires expertise in variant interpretation and segregation analysis.
- Unexpected positive findings. A pathogenic variant in a gene unrelated to the indication for testing (for example, an incidental TP53 finding on a cardiac panel that includes cancer predisposition genes) requires careful counseling about implications.
- Family planning implications. Patients considering reproductive decisions based on genetic findings need genetic counseling to understand inheritance patterns, carrier risks, and testing options such as preimplantation genetic testing.
- Pharmacogenomic complexity. When multiple pharmacogenomic variants interact to influence drug metabolism, a pharmacogenomics specialist or genetic counselor can help translate the results into prescribing guidance.
- Discordant results between clinical picture and report. When the clinical suspicion for a genetic condition is high but the report is negative, or when the report identifies a variant that does not fit the phenotype, genetics expertise can guide next steps.
- Patient or family requests for detailed explanation. Genetic counselors are trained to communicate complex genetic information in accessible language. Referring patients who want deeper understanding is appropriate and beneficial.
How AI Tools Are Changing Genomic Report Interpretation
The process of classifying variants and interpreting genomic reports has traditionally been manual and labor-intensive. A clinical geneticist or laboratory scientist reviews evidence from multiple databases (ClinVar, gnomAD, OMIM, UniProt, functional literature) and applies the ACMG/AMP criteria point by point. For a single variant, this process can take 20 to 60 minutes. For a report with multiple variants, the cumulative time is significant.
AI-powered tools are beginning to change this workflow. Platforms that integrate molecular intelligence capabilities can automate evidence gathering across databases, apply ACMG/AMP criteria with structured reasoning, and surface relevant literature, all in seconds rather than hours.
Purna AI’s Molecular Intelligence Platform (MIP), for example, performs ACMG/AMP variant classification by pulling evidence from over 30 clinical and biological databases, including ClinVar, gnomAD, OMIM, UniProt, and PharmGKB. Each classification comes with structured reasoning that shows which criteria were met and what evidence supports them, making the logic transparent and auditable. For clinicians who want to quickly check a variant’s population frequency, functional annotations, or disease associations, MIP’s natural language interface allows queries like “What is the gnomAD frequency of BRCA2 c.5946del?” or “Show me ClinVar submissions for this variant” without requiring database expertise.
This does not replace clinical judgment. The clinician still makes the final decision about patient care. But having an AI system that can rapidly synthesize evidence from dozens of sources, flag discrepancies, and present the reasoning in a structured format reduces the cognitive burden and the risk of missing relevant data.
A Practical Checklist for Reading Any Genomic Report
Use this checklist when reviewing a genomic report in your clinical practice:
1. Identify the test scope
- What type of test was performed (panel, exome, genome)?
- Which genes were included?
- What variant types can the test detect?
2. Review each variant and its classification
- Is it pathogenic, likely pathogenic, VUS, likely benign, or benign?
- Is the gene relevant to the patient’s clinical indication?
- What is the zygosity (heterozygous vs. homozygous)?
3. Map classification to action
- Pathogenic or likely pathogenic: follow gene-specific management guidelines, consider cascade testing, refer to genetic counseling.
- VUS: do not change management. Note in record. Monitor for reclassification.
- Likely benign or benign: no action needed.
4. Read the limitations section
- What can this test not detect?
- If the result is negative but clinical suspicion is high, is broader testing warranted?
5. Consider the clinical context
- Does the finding match the patient’s phenotype?
- Is the inheritance pattern consistent with the family history?
- Are there pharmacogenomic implications for current medications?
6. Decide on referral
- Does this patient need genetic counseling?
- Are the results complex enough to warrant genetics specialist review?
7. Document and follow up
- Record all findings in the medical record.
- Set a reminder to recheck VUS classifications in 1 to 2 years.
- Communicate results to the patient with appropriate context.
Bringing It Together
Reading a genomic report is a skill that every clinician ordering or receiving genetic tests should develop. The report’s structure is consistent across laboratories, the variant classification system follows established guidelines, and the rules for when to act and when to wait are well defined. The most common errors come not from the report itself but from misunderstanding what its findings mean in clinical context.
For clinicians who want to deepen their understanding of variant interpretation, the companion guide on VUS and how to resolve them provides a detailed look at the evidence types that drive reclassification. And for a broader perspective on how genomic findings translate into patient management decisions, From Variant to Visit walks through real clinical scenarios.
As genomic testing becomes standard across more areas of medicine, the ability to read and act on these reports will be as fundamental as interpreting a CBC or reading an imaging report. Building that fluency now prepares clinicians to deliver the precision medicine their patients increasingly expect.
Purna AI’s Molecular Intelligence Platform helps clinicians and researchers interpret genomic findings with ACMG/AMP-guided variant classification, evidence from 30+ databases, and structured reasoning you can audit. Apply for up to $10,000 in free research credits to try it in your workflow.
Explore Purna's Molecular Intelligence Platform
AI-powered workspace for biology teams to accelerate drug discovery from target identification to lead optimization.
Try Purna AI →
