What Is a Variant of Uncertain Significance (VUS) and How Do You Resolve One?

Every year, millions of genetic tests return at least one variant of uncertain significance. For patients, the VUS label can feel like a non-answer: something was found, but nobody can say whether it matters. For clinicians and genetic counselors, a VUS is a classification gap, a variant where the available evidence is insufficient to determine whether it causes disease or is completely benign. Resolving that gap is one of the most important challenges in clinical genomics today.
This guide explains what a variant of uncertain significance is, why so many variants receive this label, what evidence can resolve the uncertainty, and how emerging computational and structural approaches are accelerating VUS reclassification.
What Is a Variant of Uncertain Significance?
A variant of uncertain significance (VUS) is a genetic change identified through sequencing that cannot be classified as either disease-causing (pathogenic) or harmless (benign) based on the evidence currently available. It sits in the middle of the five-tier classification system defined by the American College of Medical Genetics and Genomics (ACMG) and the Association for Molecular Pathology (AMP).
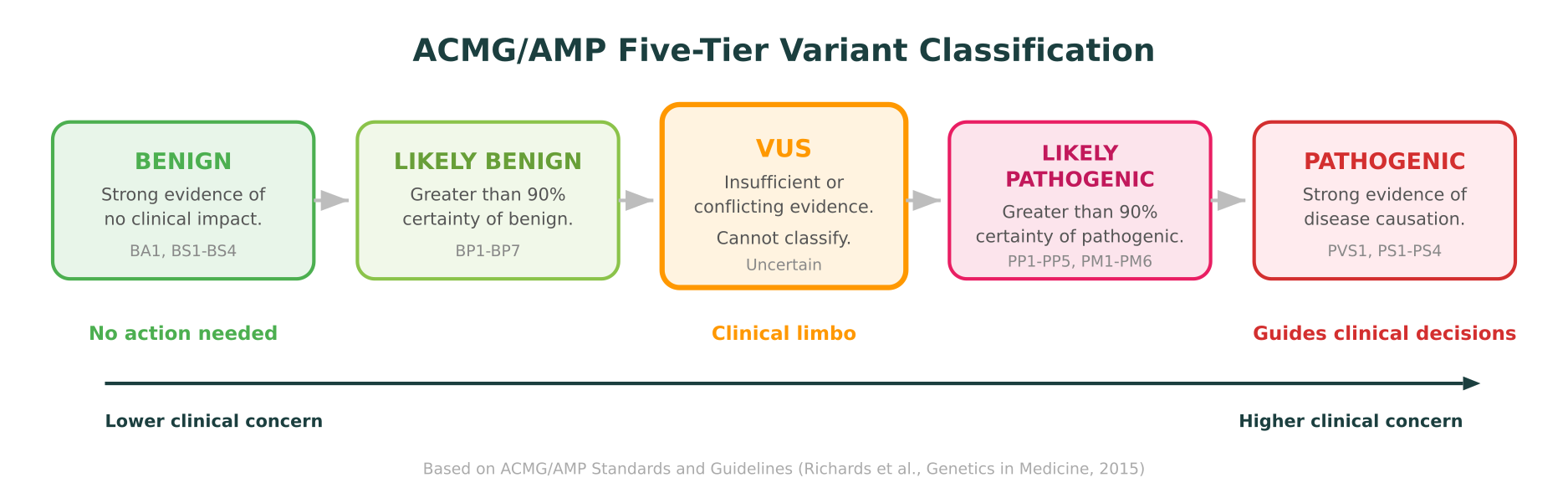
The ACMG/AMP framework, published in Genetics in Medicine in 2015 and widely adopted across clinical genetics laboratories, classifies variants into five categories:
- Pathogenic — Strong evidence that the variant causes disease.
- Likely pathogenic — Greater than 90% certainty the variant is disease-causing.
- Variant of uncertain significance (VUS) — Insufficient or conflicting evidence. Cannot be classified in either direction.
- Likely benign — Greater than 90% certainty the variant is harmless.
- Benign — Strong evidence that the variant has no clinical impact.
A VUS is not a diagnosis. It is not a confirmed finding. It is an acknowledgment that the evidence has not yet crossed the threshold needed for a definitive classification. Clinical guidelines are clear: VUS results should not be used to make medical decisions such as prophylactic surgeries, changes in screening protocols, or cascade testing of family members.
Why So Many Variants Are Classified as VUS
The VUS category is not a rare edge case. In ClinVar, the largest public repository of clinically interpreted genetic variants, roughly 50% of all unique variant submissions carry the VUS label. Several factors drive this high proportion.
Limited evidence for rare variants
Most disease-associated genes have thousands of possible missense variants, but only a fraction have been observed enough times to build a robust evidence base. A variant seen in a single patient, with no prior reports in clinical databases and no published functional studies, will almost always be classified as VUS regardless of its actual clinical impact. The evidence simply does not exist yet.
Population database gaps
Population frequency is one of the strongest signals for benign classification. If a variant is common in healthy populations (for example, allele frequency above 5% in gnomAD), it is very unlikely to cause a rare Mendelian disease. But for variants not seen or seen only once in population databases, this line of evidence is unavailable. This is particularly acute for individuals from underrepresented ancestral backgrounds, where population reference data is sparser.
A 2018 analysis in Genetics in Medicine demonstrated that individuals of African, Asian, and Latino ancestry receive VUS results at substantially higher rates than individuals of European ancestry, largely because reference databases like gnomAD have historically overrepresented European populations. While this gap has narrowed, it remains a significant equity issue in variant interpretation.
Conflicting computational predictions
In silico predictors like SIFT, PolyPhen-2, CADD, and REVEL often disagree on whether a given missense variant is damaging. When one tool predicts “tolerated” and another predicts “damaging,” the computational evidence line is inconclusive, which contributes to VUS classification rather than resolving it.
Novel variants in well-studied genes
Even genes with extensive literature, like BRCA1 or SCN5A, contain positions where specific amino acid substitutions have never been reported. A novel missense variant at a well-characterized position might have indirect evidence (other pathogenic variants at the same residue, conservation data, functional domain information), but if the specific substitution lacks direct evidence, the classification may still land at VUS.
The ACMG/AMP Evidence Framework
Understanding how to resolve a VUS requires understanding how the ACMG/AMP classification system works. The framework uses a set of evidence codes, each with a defined strength level, that are combined to reach a classification.
Evidence strength levels
The framework defines four strength tiers for evidence:
- Very strong (PVS) — e.g., predicted null variant in a gene where loss of function is a known mechanism of disease.
- Strong (PS, BS) — e.g., well-established functional studies showing a damaging or neutral effect.
- Moderate (PM) — e.g., absent from population databases, located in a mutational hotspot or functional domain.
- Supporting (PP, BP) — e.g., computational evidence, co-segregation in a single family.
To reach “likely pathogenic,” a variant typically needs at least two moderate criteria or one strong criterion plus supporting evidence. To reach “pathogenic,” combinations of strong and very strong evidence are required. The specific combination rules are codified in the 2015 ACMG/AMP guidelines and have been refined by disease-specific expert panels through ClinGen.
A VUS results when the accumulated evidence falls short of these thresholds in either direction. The variant might have one supporting criterion toward pathogenic and one toward benign, or it might simply have no evidence at all beyond being a missense change in a disease-associated gene.
What Evidence Upgrades a VUS Toward Pathogenic?
Reclassifying a VUS to likely pathogenic or pathogenic requires gathering additional evidence lines that, when combined under the ACMG/AMP rules, cross the classification threshold.
Functional studies (PS3)
Well-designed functional assays are among the strongest evidence types for reclassification. If an assay demonstrates that the variant disrupts protein function (for example, reduced enzymatic activity, impaired protein-protein interaction, or loss of channel gating), this counts as strong evidence toward pathogenicity.
The challenge is that functional assays are expensive, time-consuming, and gene-specific. Multiplexed assays of variant effect (MAVEs), sometimes called deep mutational scanning, are beginning to change this. Large-scale MAVE datasets now exist for genes like BRCA1 (Nature, 2018), TP53 (Nature Genetics, 2023), and MSH2 (Nature Genetics, 2024). These datasets provide functional evidence for thousands of variants simultaneously, enabling reclassification at scale.
Segregation data (PP1)
If a variant co-segregates with disease in multiple affected family members (and is absent in unaffected members), this provides evidence toward pathogenicity. The strength of the evidence increases with the number of informative meioses. Segregation in a single small family provides supporting evidence. Segregation across multiple large families can reach moderate or strong evidence levels.
De novo occurrence (PS2)
If the variant occurred de novo (present in the proband but absent in both biological parents), and the patient’s phenotype matches the expected disease for that gene, this is strong evidence toward pathogenicity. Confirmed de novo status requires testing of both parents, which is not always performed.
Computational and structural evidence (PP3)
Computational predictions, when multiple tools agree that a variant is damaging, contribute supporting evidence. This includes sequence-based predictors (REVEL, CADD) as well as structural evidence: protein instability predicted by tools like DynaMut2, high evolutionary conservation at the affected residue, and location within a functional domain or active site.
Structural evidence is especially valuable for missense variants where the amino acid change affects the protein’s three-dimensional fold. A variant that introduces a large, charged residue into a tightly packed hydrophobic core is far more likely to be destabilizing than one at a flexible, surface-exposed position. This type of reasoning requires protein structure analysis, not just sequence-level prediction.
For a detailed walkthrough of how protein structure analysis contributes to variant interpretation, see How to Predict the Impact of Protein Mutations on Structure and Function.
Population rarity (PM2)
Absence or extreme rarity in population databases like gnomAD provides moderate supporting evidence. If a variant has never been observed in over 800,000 alleles across diverse populations, it is less likely to be a benign polymorphism. However, rarity alone does not prove pathogenicity, as most rare variants are still benign.
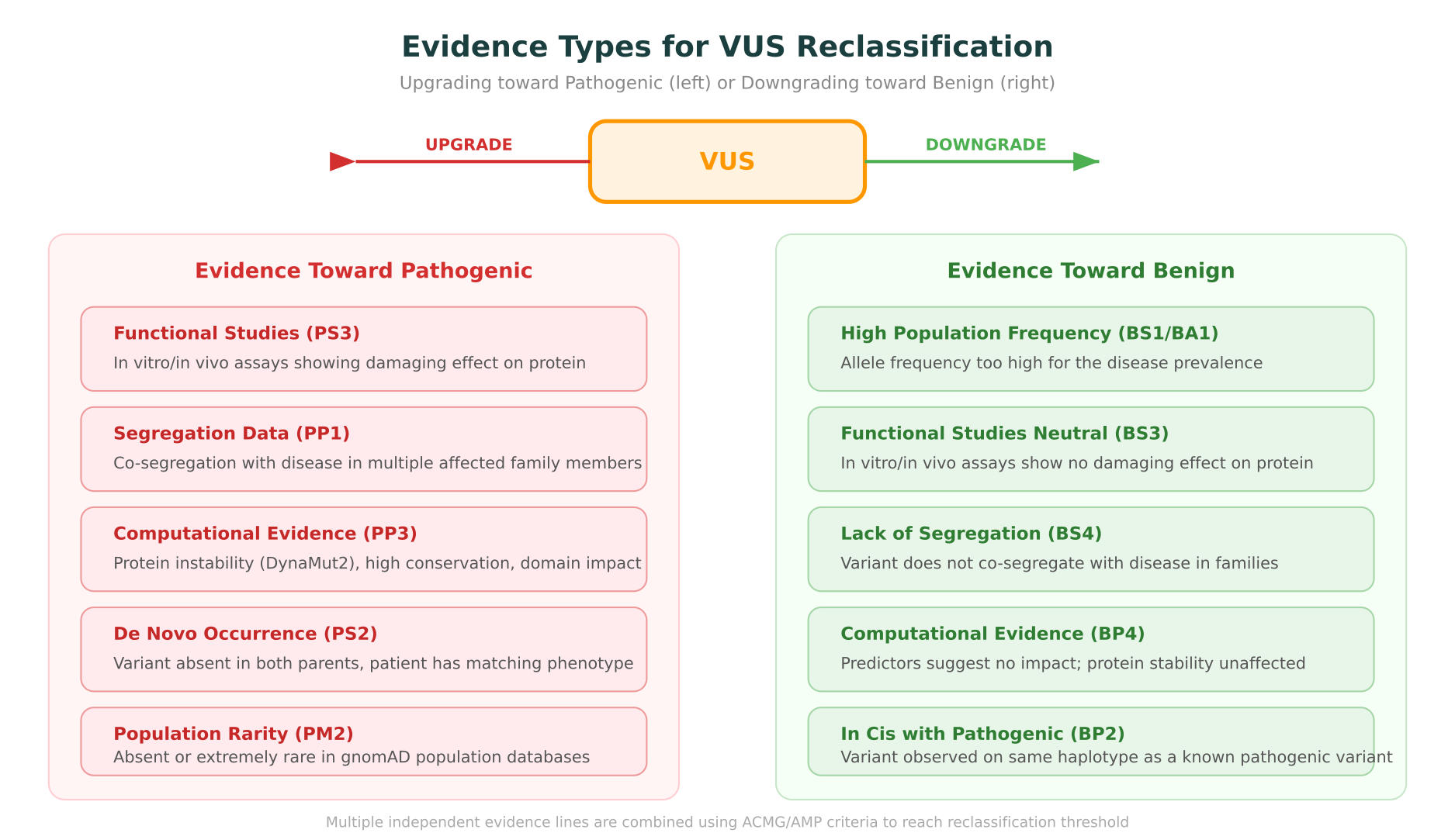
What Evidence Downgrades a VUS Toward Benign?
The same framework works in the opposite direction. Evidence that a variant is tolerated or common pushes classification toward likely benign or benign.
High population frequency (BS1, BA1)
If a variant is observed at a frequency higher than expected for the disease prevalence, it is unlikely to be pathogenic. The BA1 stand-alone benign criterion applies to variants with allele frequency above 5% in any general continental population in gnomAD. Disease-specific frequency thresholds, often much lower, are defined by ClinGen expert panels.
Functional studies showing no effect (BS3)
If a well-validated functional assay shows that the variant does not impair protein function, this counts as strong benign evidence. MAVE datasets are particularly useful here because they test both damaging and neutral effects across thousands of positions.
Lack of segregation (BS4)
If the variant is present in unaffected individuals within a family, or does not co-segregate with the disease, this is evidence against pathogenicity.
Computational evidence for benign (BP4)
When multiple in silico predictors agree that a variant is tolerated, and structural analysis shows no effect on protein stability or function, this provides supporting benign evidence.
Observation in healthy individuals (BS2)
If the variant has been observed in well-phenotyped healthy individuals at an age when the disease would be expected to manifest, this is strong benign evidence.
How Long Does VUS Reclassification Take?
Reclassification is not instant. The timeline varies dramatically depending on the gene, the variant, and the pace at which new evidence accumulates.
A large-scale study of ClinVar reclassification rates published in Genetics in Medicine in 2022 found that approximately 10% of VUS variants are reclassified within five years, and the majority of reclassifications move toward benign rather than pathogenic. This asymmetry makes sense: as population databases grow, many rare variants are eventually found at higher frequencies, pushing them toward benign.
For clinically actionable genes with active research communities (like BRCA1, BRCA2, KCNQ1, and SCN5A), reclassification rates are faster because new evidence (functional studies, case reports, population data) accumulates more quickly. For less-studied genes, variants can remain classified as VUS for a decade or longer.
Several factors are accelerating reclassification timelines:
- Growing population databases. gnomAD v4, released in late 2023, expanded to over 800,000 exomes and 76,000 genomes, providing frequency data for variants that were previously unobserved.
- Multiplexed functional assays. MAVE datasets provide functional evidence for thousands of variants at once, bypassing the bottleneck of one-at-a-time functional studies.
- ClinGen expert panels. Gene-specific variant curation expert panels (VCEPs) systematically review and reclassify variants using standardized, adapted ACMG criteria.
- Data sharing initiatives. Platforms like ClinVar, LOVD, and GeneMatcher enable laboratories and clinicians to share variant observations, increasing the evidence base for rare variants.
How Structural and Computational Evidence Helps Resolve VUS
For missense variants, where a single amino acid is substituted for another, structural and computational evidence can provide classification-relevant information even when clinical data is limited.
Protein stability analysis
Tools like DynaMut2 predict the change in thermodynamic stability (delta-delta-G) caused by a mutation. A variant that significantly destabilizes the protein fold (large negative DDG) is more likely to impair function. This evidence contributes to the PP3 (computational evidence supports a deleterious effect) criterion under ACMG/AMP guidelines.
Stability analysis is most informative when combined with structural context. A destabilizing mutation in a buried residue within a structured domain is a stronger signal than the same DDG value at a surface-exposed, flexible position. This is why integrating structure retrieval, visualization, and stability prediction into a single workflow matters.
Conservation analysis
Evolutionary conservation at the mutation site provides independent evidence. If the affected residue is highly conserved across species, substitutions at that position are more likely to be functionally important. Tools like ConSurf calculate per-residue conservation scores from multiple sequence alignments, and AlphaMissense (published in Science, 2023) provides pathogenicity scores for all possible human missense variants based on evolutionary and structural features.
Functional domain and active site proximity
ACMG criterion PM1 applies when a variant falls in a mutational hotspot or well-established functional domain without benign variation. Knowing whether a variant sits in a catalytic active site, a protein-protein interaction interface, or a critical structural element requires protein structure analysis, either from experimentally determined structures in the PDB or predicted structures from AlphaFold.
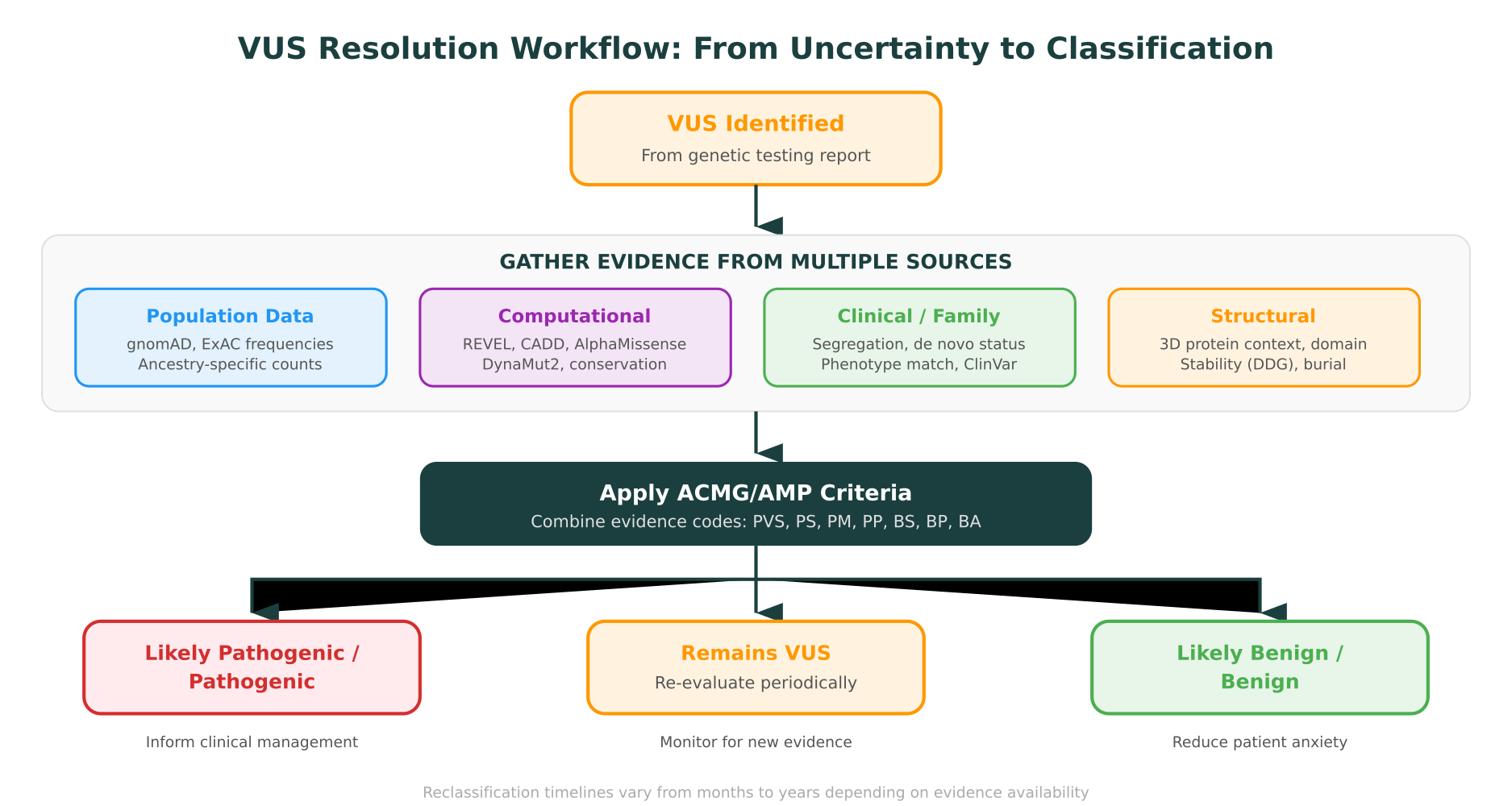
Integrating multiple computational lines
No single computational tool provides definitive evidence for variant classification. The power comes from integrating multiple lines: if a variant is predicted destabilizing by DynaMut2, highly conserved by ConSurf, classified as likely pathogenic by AlphaMissense, located in a functional domain (PM1), and absent from population databases (PM2), the combined evidence may be sufficient to upgrade from VUS to likely pathogenic.
This kind of multi-evidence synthesis is precisely what molecular intelligence platforms are designed to do. Rather than running each analysis in a separate tool and manually combining results, an integrated platform can pull structure, predict stability, check conservation, query ClinVar and gnomAD, and synthesize the evidence into a structured interpretation with citations.
The Role of AI in Accelerating VUS Resolution
The bottleneck in VUS resolution has historically been evidence gathering: manually searching databases, running individual computational tools, synthesizing results across platforms. AI-powered approaches are compressing this process.
AI-powered variant interpretation
Platforms that integrate AI with biological databases can automate much of the evidence-gathering workflow for VUS resolution. Instead of a geneticist manually querying ClinVar, gnomAD, OMIM, UniProt, and a protein structure database in separate tabs, an AI system can retrieve all relevant evidence in response to a single query, apply ACMG/AMP criteria systematically, and generate a structured interpretation showing which criteria are met and which are not.
This does not replace expert review. The ACMG/AMP guidelines require professional judgment, particularly for criteria that involve clinical correlation (phenotype matching, family history assessment, consideration of genetic heterogeneity). But the time spent on evidence retrieval and initial classification can be reduced from hours to minutes.
Structural AI for computational evidence
The rise of AI-predicted protein structures, particularly from AlphaFold and related tools, has expanded the applicability of structural evidence to variants in proteins that lack experimental structures. Previously, structural evidence was only available for the subset of proteins with crystal structures in the PDB. Now, predicted structures cover nearly every human protein, enabling stability analysis and domain assessment for a much broader set of variants.
For a comparison of protein structure prediction tools and how they apply to variant analysis, see AlphaFold 3 vs Boltz-2 vs ESMFold.
Large-scale computational reclassification
Several research groups have demonstrated AI-assisted reclassification of VUS at scale. By systematically applying computational predictions, population frequency filters, and structural analyses to large sets of ClinVar VUS, these approaches identify subsets of variants with strong computational evidence toward benign or pathogenic classification. While computational evidence alone rarely crosses the ACMG threshold for definitive reclassification, it can prioritize which variants should receive further laboratory or clinical follow-up.
Clinical Implications of VUS for Patients and Providers
The clinical management of a VUS finding requires careful communication and appropriate follow-up.
What a VUS means for patients
Receiving a VUS result is understandably frustrating. Patients often seek genetic testing for definitive answers, and a VUS provides the opposite. Genetic counselors play a critical role in explaining that a VUS is not a positive finding, should not drive clinical decisions, and may be reclassified over time as new evidence emerges.
Key points for patient communication:
- A VUS is not a diagnosis. It does not confirm a genetic condition.
- Clinical management should be guided by personal and family history, not by the VUS result.
- The variant may be reclassified in the future. Periodic re-contact or re-analysis is recommended.
- Family members generally should not be tested for the specific VUS (cascade testing is not recommended for VUS).
Periodic re-analysis
Variant reclassification is an ongoing process. Clinical laboratories periodically re-evaluate VUS when new evidence becomes available, and patients can request re-analysis of their results. The frequency and process for re-analysis varies by laboratory and healthcare system, but intervals of one to three years are common for high-interest genes.
The clinical genetics community has increasingly advocated for systematic re-analysis programs. A study in Genetics in Medicine (2019) found that systematic annual re-analysis of VUS yielded clinically significant reclassifications in approximately 7% of cases per year, primarily in genes with active research and growing population data.
The management gap
The practical challenge is that variant-to-visit workflows depend on clear classifications. When a variant is classified as VUS, the typical recommendation is to follow clinical phenotype, not genetics. This means patients with a VUS in a cancer predisposition gene like BRCA2 follow standard screening guidelines rather than the enhanced screening recommended for known pathogenic variant carriers.
This approach is clinically appropriate but creates a management gap: some proportion of VUS variants are genuinely pathogenic, and the patients carrying them are not receiving the intensified management they might benefit from. Accelerating reclassification directly reduces this gap.
How MIP Supports VUS Resolution
Purna AI’s Molecular Intelligence Platform (MIP) integrates the key evidence sources and analysis tools needed for VUS interpretation into a single workspace. Rather than switching between ClinVar, gnomAD, UniProt, PDB, and separate computational tools, researchers and clinicians can query a variant and receive a structured interpretation that includes:
- ACMG/AMP classification with structured reasoning. MIP applies classification criteria systematically, showing which evidence codes are met, which are not, and citing the source databases for each line of evidence.
- Protein structure analysis. MIP retrieves structures from PDB or AlphaFold, renders them in Molstar for visualization, and runs DynaMut2 for stability predictions, all within the same workspace.
- 30+ database integrations. ClinVar, gnomAD, OMIM, UniProt, PDB, PharmGKB, LOVD, and more. Every piece of evidence is traceable to its source.
- Natural language interface. A clinician can ask “What is the evidence for or against pathogenicity of NM_000059.4:c.7397T>C in BRCA2?” and receive a synthesized response covering population frequency, computational predictions, structural context, ClinVar submissions, and functional domain analysis.
This approach does not bypass the need for expert judgment. It compresses the evidence-gathering phase so that the expert’s time is spent on interpretation and clinical correlation rather than data retrieval.
Looking Ahead: Reducing the VUS Burden
The proportion of VUS results will continue to decline as population databases expand, functional assay datasets grow, and AI-powered interpretation tools mature. But for the foreseeable future, VUS will remain a significant fraction of clinical genetic test results, and the need for systematic, evidence-based resolution workflows will persist.
The most impactful advances are likely to come from three directions:
- Broader population representation in reference databases. Reducing the disparity in VUS rates across ancestral backgrounds requires more diverse population sequencing data.
- Scaled functional evidence. Multiplexed assays of variant effect for clinically important genes will provide functional data for thousands of variants that currently lack it.
- Integrated computational platforms. Tools that combine population, computational, structural, and clinical evidence into a single assessment, with transparent reasoning and citations, will make VUS resolution more systematic and less dependent on individual expertise.
The goal is not to eliminate uncertainty. Some variants will remain genuinely uncertain, particularly novel variants in poorly characterized genes. The goal is to ensure that every available line of evidence is efficiently gathered, correctly weighted, and transparently presented so that the classification reflects the current state of knowledge.
For researchers and clinicians working through VUS interpretation today, molecular intelligence provides the infrastructure to accelerate that process: evidence synthesis across databases, computational analysis in one workspace, and structured reasoning traceable to source data.
Explore how MIP can support your variant interpretation workflows at purna.ai.
MIP is Purna AI’s Molecular Intelligence Platform, an AI-powered workspace for biology teams. ACMG/AMP variant classification with structured reasoning, protein structure prediction and visualization, DynaMut2 stability analysis, and 30+ database integrations in one place. Explore the platform at purna.ai. Researchers can apply for up to $10,000 in free credits to run their analyses on MIP.
Explore Purna's Molecular Intelligence Platform
AI-powered workspace for biology teams to accelerate drug discovery from target identification to lead optimization.
Try Purna AI →
