Best Bioinformatics Platforms in 2026: Galaxy, Terra, and Cloud Alternatives Compared

Choosing the best bioinformatics platform in 2026 is a fundamentally different exercise than it was even two years ago. The question has shifted from “which pipeline runner should my team use?” to “which intelligence layer do we need on top of our data?” Traditional platforms like Galaxy and Terra remain indispensable for specific use cases, but a new category of AI-native biology platforms has entered the market, and they are changing what researchers expect from their computational infrastructure.
This comparison covers nine major platforms across three categories: open-source workflow tools, enterprise compliance platforms, and AI-native biology platforms. We evaluate each on pricing, learning curve, pipeline support, database integrations, AI capabilities, collaboration features, and who each platform is best suited for. The goal is to help lab leads, bioinformatics managers, and research directors make an informed decision based on their team’s actual needs.
If you are weighing the broader open-source versus commercial decision, our detailed breakdown in Open-Source vs Commercial Bioinformatics Platforms covers the cost, compliance, and operational tradeoffs in depth.
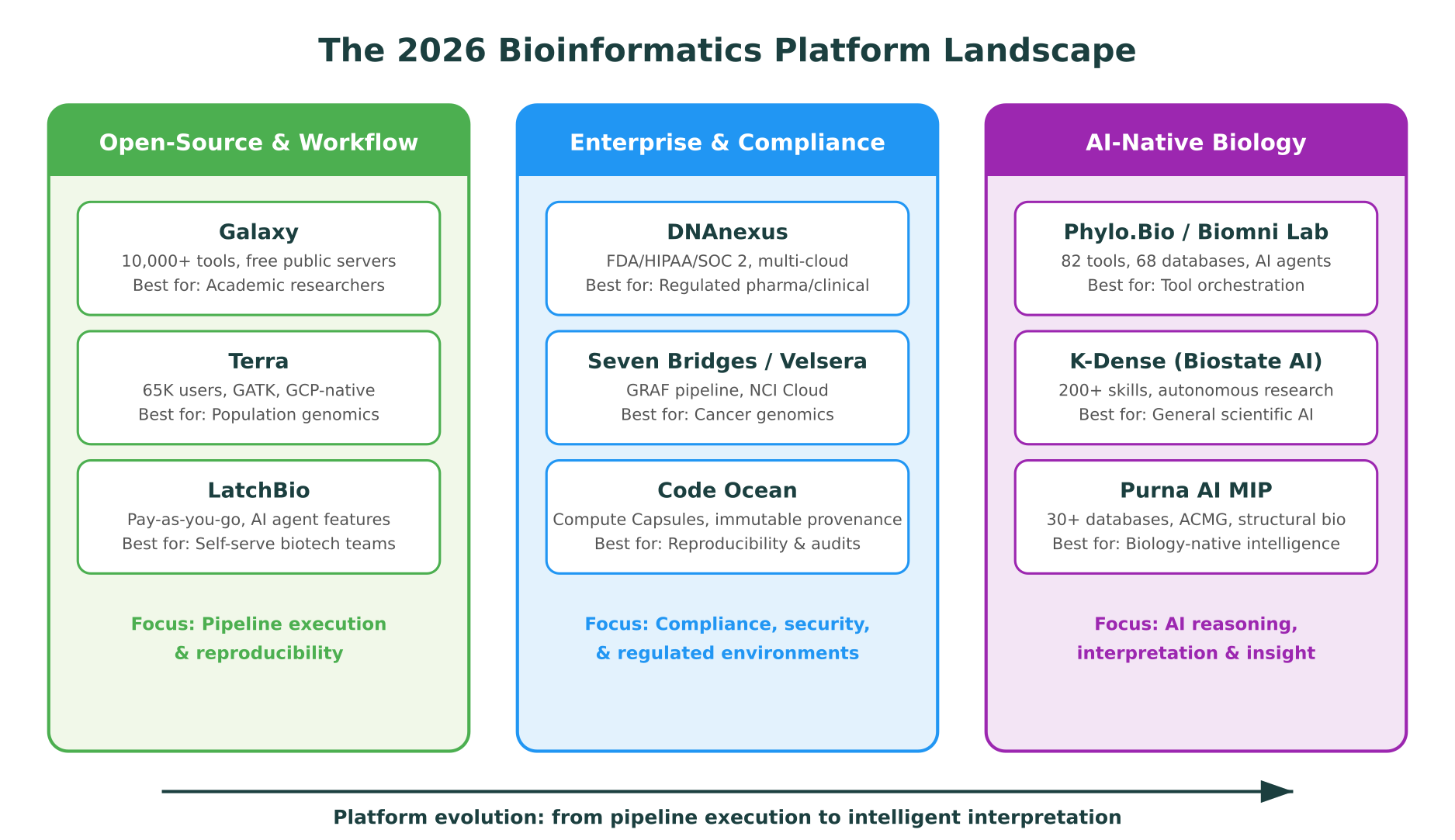
Open-Source and Workflow Platforms
These platforms focus on pipeline execution, reproducibility, and giving researchers access to established bioinformatics tools. They have the largest user bases and the deepest community support. For teams with bioinformatics expertise, they remain the backbone of computational biology.
Galaxy: the community-driven workbench
Galaxy has been the entry point for accessible bioinformatics since 2005. It provides a web-based graphical interface for building and running analysis workflows without writing code. The public Galaxy servers (usegalaxy.org, usegalaxy.eu, usegalaxy.org.au) offer free access to over 10,000 tools covering genomics, proteomics, metabolomics, and more.
Strengths. Galaxy’s greatest asset is its ecosystem. The Galaxy Training Network provides hundreds of tutorials across dozens of disciplines. The community is global, active, and welcoming to newcomers. For academic researchers running standard pipelines (RNA-seq, variant calling, ChIP-seq, metagenomics), Galaxy’s public servers provide a genuine zero-cost starting point. Workflow reproducibility is built in, with every analysis tracked and shareable.
Limitations. Galaxy’s web interface, while more accessible than the command line, shows its age. Complex multi-step analyses require significant understanding of tool parameters and data formats. Public servers impose resource limits that make large-scale analyses impractical. Running your own Galaxy instance requires server administration, tool installation, and ongoing maintenance. And critically, Galaxy has no AI reasoning layer. It executes tools but does not interpret results, connect findings across databases, or provide biological context. For teams that need bioinformatics without coding expertise, the learning curve is still substantial despite the GUI.
Pricing. Free on public servers. Self-hosted costs vary by infrastructure (typically $20,000 to $50,000 per year for a well-maintained institutional instance including cloud compute).
Best for. Academic researchers who need reproducible workflows, have some computational experience, and work within the resource limits of public servers.
Terra: the genomics gold standard
Terra, developed by the Broad Institute and Verily, is the cloud-native platform that much of the large-scale genomics community relies on. Built on Google Cloud Platform (with Azure support expanding), Terra provides managed infrastructure for running WDL (Workflow Description Language) pipelines, Jupyter and R notebooks, and accessing major public datasets.
The numbers speak to its scale: over 65,000 registered users, more than 80 petabytes of data under management, and over 30 million workflows executed. Terra is tightly integrated with GATK best practices pipelines, and its data library includes flagship datasets like gnomAD, TOPMed, and the All of Us Research Program.
Strengths. For population-scale genomics, Terra is the standard. The integration with GATK means that germline variant calling, somatic mutation detection, and joint genotyping workflows are well-tested and documented. The platform handles cloud provisioning, auto-scaling, and cost optimization. There is no platform fee; you pay only for the GCP compute and storage you consume. The open-source nature of Terra (built on Cromwell for workflow execution) means you are not locked into a black-box system.
Limitations. Terra’s learning curve is steeper than newcomers expect. Configuring workspaces, writing or modifying WDL workflows, understanding data models, and managing GCP costs all require technical sophistication. The platform is primarily tied to Google Cloud, which can be a constraint for institutions with multi-cloud mandates. And while Terra excels at pipeline execution, it does not provide AI-assisted analysis. You can run a GATK pipeline to produce a VCF file, but interpreting those variants, connecting them to clinical databases, and generating a report requires additional tools or manual effort.
Pricing. No platform fee. GCP compute and storage billed directly (a whole-genome analysis typically costs $3 to $5 per sample for standard pipelines).
Best for. Large consortia, population genomics projects, GATK users, and teams with dedicated bioinformatics staff comfortable with WDL and cloud infrastructure.
LatchBio: self-serve cloud bioinformatics
LatchBio occupies the middle ground between traditional pipeline platforms and the newer AI-native tools. It provides a cloud bioinformatics environment with a more modern interface than Galaxy or Terra, and has recently begun adding AI agent features (built on Claude) for natural language interaction.
Strengths. LatchBio’s pricing model is refreshingly transparent: 1 credit equals $1, with CPU compute at approximately $0.003 per core per minute and GPU instances ranging from $0.52 to $4.83 per hour. There is no platform fee, no lock-in, and no minimum commitment. The interface is designed for biologists, not just bioinformaticians, and the growing AI agent capabilities suggest the platform is moving toward natural language workflows. GPU access makes it viable for deep learning workloads alongside traditional bioinformatics.
Limitations. LatchBio’s community is smaller than Galaxy’s or Terra’s. The AI features are new and still maturing. The platform lacks the depth of specialized compliance certifications that regulated environments require. And while the direction is promising, it does not yet offer the biology-specific intelligence (variant classification, structural analysis, multi-database reasoning) found in purpose-built AI biology platforms.
Pricing. Pay-as-you-go. CPU ~$0.003/core/min. GPU $0.52 to $4.83/hr depending on instance type. No platform fee.
Best for. Biotech teams that want self-serve bioinformatics without enterprise pricing, especially those looking for a modern interface and emerging AI capabilities.
Enterprise and Compliance Platforms
For organizations operating under regulatory oversight (pharma companies, clinical labs, government agencies), compliance is not a feature; it is a requirement. These platforms prioritize security certifications, audit trails, and validated environments.
DNAnexus: industry-leading compliance
DNAnexus has established itself as the platform of choice for regulated bioinformatics environments. It supports FDA 21 CFR Part 11 compliance, HIPAA, SOC 2, and GDPR, with multi-cloud deployment across AWS, Azure, and private cloud infrastructure. The Apollo platform provides data management capabilities purpose-built for genomic data governance.
In September 2025, DNAnexus announced a partnership with Oracle to deliver genomic insights at the point of care, signaling the platform’s push deeper into clinical workflows.
Strengths. DNAnexus has the most comprehensive compliance portfolio in the bioinformatics platform market. Pharma companies, clinical diagnostic labs, and government health agencies trust it for good reason. Multi-cloud support means organizations are not locked into a single provider. The API and app framework allow custom analysis development, and enterprise support tiers ensure responsive help.
Limitations. DNAnexus is priced for enterprise customers. There is no free tier, and academic labs typically find the licensing costs prohibitive. Despite the polished infrastructure, building custom analyses still requires bioinformatics programming skills. And like Terra, DNAnexus is a pipeline execution and data management platform. It does not provide AI reasoning, variant interpretation, or cross-database synthesis.
Pricing. Enterprise licensing (custom quotes). No free tier.
Best for. Pharma companies, clinical diagnostic labs, and government agencies requiring validated, compliant genomic data platforms.
Seven Bridges / Velsera: cancer genomics depth
Seven Bridges, now operating under the Velsera umbrella, brings particular strength in oncology and precision medicine. Their GRAF pipeline uses graph-based variant calling to reduce reference bias, a meaningful advancement for population genetics. The ARIA platform supports multi-omics analysis, and their infrastructure powers the NCI Cancer Genomics Cloud and CAVATICA for pediatric genomics research.
Strengths. Graph-based variant calling through GRAF addresses a real limitation of linear reference genomes: population bias in variant detection. The NCI partnership gives Seven Bridges unmatched access to cancer genomic datasets. For oncology research teams, the combination of graph-based calling, NCI data access, and compliance certifications is a strong value proposition.
Limitations. The platform has a steep learning curve, with CWL (Common Workflow Language) as the primary workflow language. The focus on cancer and precision medicine, while deep, means the platform is narrower than more general-purpose alternatives. Pricing is enterprise-oriented and not publicly available.
Pricing. Enterprise licensing (custom quotes).
Best for. Oncology research teams, cancer genomics centers, and organizations working with NCI datasets.
Code Ocean: reproducibility as infrastructure
Code Ocean takes a different approach from the pipeline-focused platforms. It is fundamentally a reproducibility platform, built around the concept of Compute Capsules: self-contained, executable research environments that capture code, data, environment, and results in an immutable, citable unit.
Strengths. Code Ocean’s Lineage Graph tracks the complete provenance of every analysis, from raw data through every transformation to final results. MLflow integration supports machine learning model tracking. The platform imports nf-core pipelines and supports multiple languages and frameworks. For organizations that need audit trails, immutable provenance, and the ability to reproduce any analysis months or years later, Code Ocean is best in class.
Limitations. Code Ocean is not biology-specific. It provides no biological intelligence, no variant classification, no database integrations. It is an excellent general-purpose reproducibility platform that happens to work well for bioinformatics. Pricing is not publicly available and is oriented toward institutional licenses.
Pricing. Institutional licensing (custom quotes).
Best for. Research organizations, journals, and regulatory bodies that need computational reproducibility and audit trails.
AI-Native Biology Platforms: The New Category
This is where the bioinformatics landscape is shifting most rapidly. A new generation of platforms treats AI not as an add-on but as the core infrastructure for biological analysis. Rather than asking “how do I run this pipeline?”, researchers can ask “what does this variant mean?” or “which proteins are most affected by this mutation?” and receive evidence-backed answers.
Three platforms define this emerging category, each with a distinct approach.
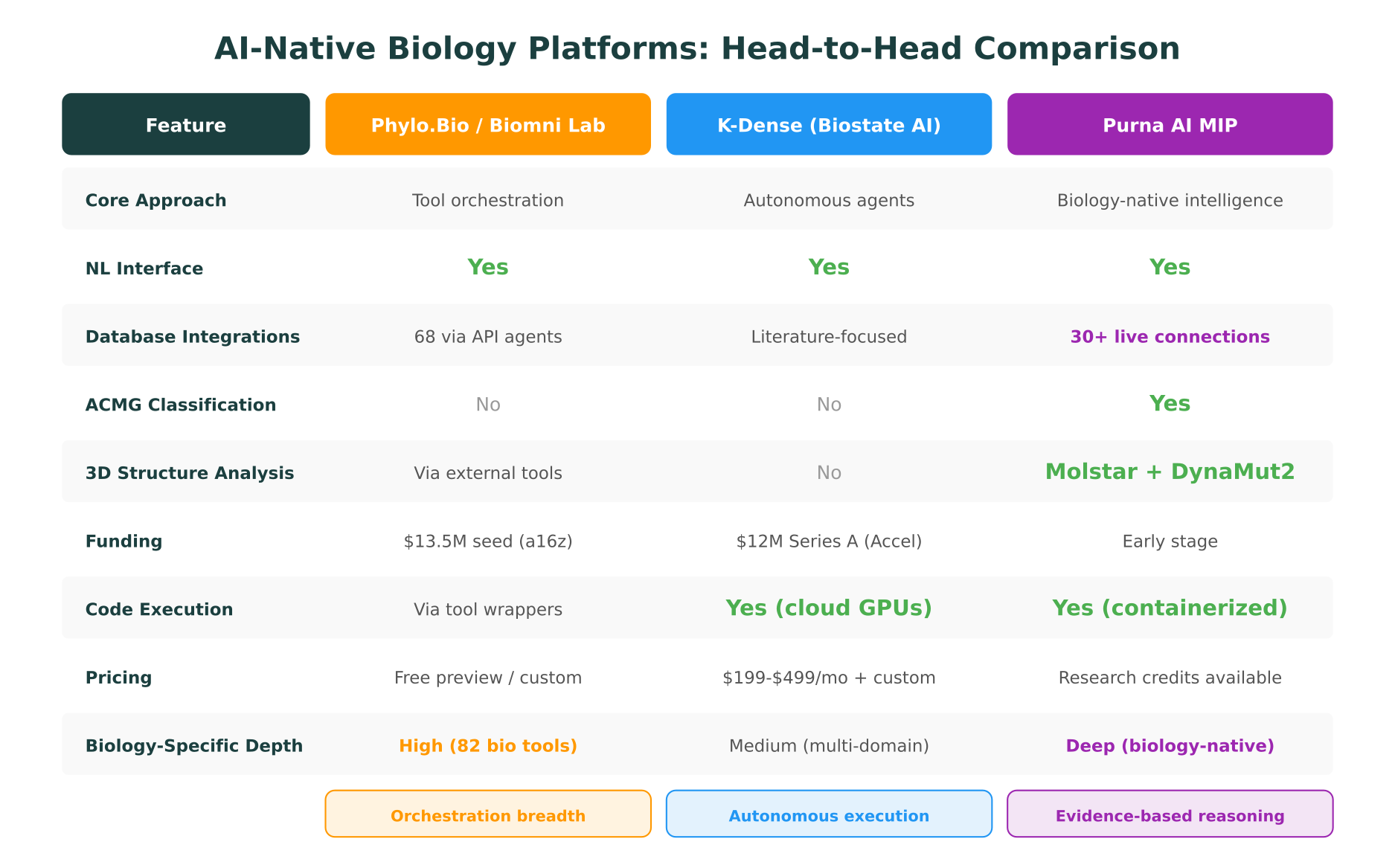
Phylo.Bio / Biomni Lab: tool orchestration at scale
Phylo.Bio is a Stanford spin-out founded by Kexin Huang and Jure Leskovec, with an advisory board that includes Nobel Laureate Carolyn Bertozzi and Feng Zhang. The platform, which launched its research preview in February 2026, calls itself an “Integrated Biology Environment” (IBE) and uses the “IDE for Biology” framing to describe its vision.
The platform deploys AI agents that orchestrate 82 biology tools, query 68 databases, and coordinate over 100 software packages. The interface is conversational, allowing researchers to describe analyses in natural language. Phylo.Bio is built on the open-source Biomni project, which reports adoption by over 7,000 labs, including 18 of the top 20 pharmaceutical companies.
Strengths. The orchestration breadth is impressive. Ginkgo Bioworks published a case study describing analyses that previously took weeks being completed in hours using Phylo.Bio’s agent system. The open-source Biomni foundation gives the platform transparency and community engagement. The advisory and investor pedigree is among the strongest in the space.
Limitations. Phylo.Bio launched its research preview in February 2026 and does not yet have an enterprise track record. The platform orchestrates existing tools via AI agents rather than providing native biological intelligence. This means it routes queries to the right tool and returns results, but the depth of reasoning within each biological domain (variant interpretation, structural analysis, multi-omics correlation) depends on the underlying tools being orchestrated. Quality during research preview will differ from production readiness.
Pricing. Free during research preview. Enterprise pricing is custom and not yet publicly available.
Best for. Research labs that want a conversational interface to existing bioinformatics tools and value orchestration breadth over domain-specific depth.
K-Dense by Biostate AI: autonomous scientific agents
K-Dense, developed by Biostate AI, takes the most ambitious approach in the AI-native category. It positions itself as an autonomous AI agent platform for scientific research, deploying multi-agent systems that handle planning, literature review, code execution, analysis, and report generation.
The platform reports over 200 scientific skills and publishes BixBench benchmark results that show strong performance: 29.2% on BixBench, compared to GPT-5’s 22.9% and Claude 3.5 Sonnet’s 18%. K-Dense has been validated in collaboration with Harvard Medical School researcher David Sinclair on longevity research applications.
Funding includes a $12 million Series A led by Accel, with individual investors including Dario Amodei (CEO of Anthropic), Emily Leproust (CEO of Twist Bioscience), and Mike Schnall-Levin (CSO of 10x Genomics).
Strengths. The benchmark results are noteworthy. The Harvard Medical School validation provides credibility. The pricing is transparent: Plus at $199 per month (300 credits), Team at $499 per month, and custom Enterprise tiers. The multi-agent architecture that handles end-to-end research workflows, from literature review to code execution to report generation, is the most complete autonomous research vision in the market.
Limitations. K-Dense is not biology-specific. It covers finance, engineering, and other scientific domains alongside biology. This breadth means the platform’s biological depth may not match specialized tools for domain-specific tasks like ACMG variant classification or protein stability prediction. The team is currently very small (three people), which raises questions about support capacity and development velocity. And as with any autonomous AI system, users need to carefully validate outputs.
Pricing. Plus: $199/month (300 credits). Team: $499/month. Enterprise: custom.
Best for. Research teams that want autonomous end-to-end scientific workflows and are comfortable with a general-purpose AI agent that covers biology among other domains.
Purna AI MIP: biology-native intelligence
Purna AI’s Molecular Intelligence Platform (MIP) takes a different approach from both the orchestration model and the general-purpose agent model. MIP is built specifically for biology teams, with AI reasoning that is native to biological workflows rather than bolted on or generalized from other domains.
The platform connects to over 30 clinical and biological databases in real time, including ClinVar, gnomAD, OMIM, UniProt, PDB, PharmGKB, and LOVD. When you ask MIP about a variant, it does not search cached training data. It queries live databases, and synthesizes evidence across sources.
MIP has several in-chat capabilities like 3D Protein Structure Prediction, Comparing Sequence Alignments, Creating Detailed Charts, and much more. This integration means the path from “I found a variant” to “here is its structural impact” happens within a single workspace.
The platform also supports containerized code execution (Python, R, and standard bioinformatics packages), multi-omics analysis across genomics, transcriptomics, proteomics, and metabolomics, and a natural language interface for querying databases that returns cited results rather than hallucinated summaries. This concept of molecular intelligence, where AI serves as the connective layer across biological data types, represents a shift from tool-based workflows to intelligence-based workflows. These long-running jobs can write pipelines, refine them, and work without constant babysitting.
MIP provides a Parallel Hypothesis Exploration Mode where researchers can let different personalities of agents debate with each other and come back to strong conclusions with citations.
MIP also has a very strong genetic workbench platform where you can upload an unannotated VCF and find useful information about the case. You can tag a case and run queries on the case using natural language in Chat Mode. It also has several important features built for Germline cases like Pathogenicity Predictor, Notes, Population Frequency, ACMG Classigication, AI Summary, Clinical Whitelabeled Reporting, and more.
Strengths. MIP’s differentiator is depth over breadth. Rather than orchestrating many tools loosely, MIP provides native intelligence within the biological domains it covers. ACMG classification is built in, not farmed out to a third-party tool. Structural analysis is integrated, not a separate workflow. Every answer comes with citations to source databases, allowing researchers to verify the evidence chain. The combination of variant interpretation, structural biology, code execution, and multi-omics in a single workspace means fewer context switches and faster time to insight.
Limitations. MIP is an early-stage platform. Its depth in variant interpretation and structural biology is strong, but teams with very specialized needs (say, graph-based variant calling for diverse populations or large-scale population genomics) may still need domain-specific tools like Seven Bridges or Terra alongside MIP. The platform is focused on biology teams and is not a general-purpose scientific AI.
Pricing. Starts at $20/Month and $50/Month for Pro Usage.
Best for. Biology teams that need an integrated workspace combining variant interpretation, structural biology, multi-omics analysis, and code execution with AI reasoning grounded in live database evidence whether or not they are using any other tool.
How the three AI-native platforms differ
The distinction matters. Phylo.Bio is an orchestration layer: it connects existing tools through AI agents and coordinates them via natural language. This gives breadth but depends on the underlying tools for depth. K-Dense is an autonomous research agent: it handles end-to-end scientific workflows but spans multiple domains, meaning biology is one of many areas rather than the sole focus. MIP is biology-native intelligence: the AI reasoning is purpose-built for biological questions, with ACMG classification, structural analysis, and multi-omics interpretation as first-class capabilities rather than integrations.
All three represent meaningful progress beyond traditional pipeline platforms. The right choice depends on whether your team needs breadth of tool access (Phylo.Bio), autonomous end-to-end research execution (K-Dense), or depth of biology-specific reasoning (MIP).
Emerging Players Worth Watching
The bioinformatics platform market is seeing rapid entry from specialized startups. Several are worth noting even if they are too early for a full evaluation.
Bionl (bionl.ai) focuses specifically on clinical genetic testing, with an ACMG-compliant variant AI classifier (VAIC) and reports over 8,000 users. For clinical genetics labs, this focused approach may complement broader platforms.
Attma (attma.bio) positions itself as an “AI Bioinformatician” with natural language interfaces for RNA-seq and single-cell analysis. The platform is aimed at wet-lab scientists who need bioinformatics results without the computational overhead.
Benchling remains the dominant player in lab informatics (electronic lab notebooks, LIMS, molecular biology tools) but does not compete directly in the computational bioinformatics space. Many labs use Benchling alongside a bioinformatics platform.
Geneious provides desktop-based molecular biology tools for sequence analysis, alignment, and phylogenetics. It is well-established for molecular biology workflows but does not extend into the AI-native or cloud-scale analysis categories.
Comprehensive Comparison Table
| Platform | Type | AI / NL Interface | Database Integrations | Compliance | Pricing | Best For |
|---|---|---|---|---|---|---|
| Galaxy | Open-source workflow | No | Manual | User responsibility | Free (public servers) | Academic researchers |
| Terra | Cloud genomics | No | GATK/public datasets | GCP security | Pay-per-compute (no platform fee) | Population genomics, GATK users |
| LatchBio | Cloud bioinformatics | Emerging (Claude-based) | Limited | Basic | Pay-as-you-go ($0.003/core/min) | Biotech self-serve |
| DNAnexus | Enterprise genomics | No | Apollo data management | FDA/HIPAA/SOC 2/GDPR | Enterprise licensing | Regulated pharma/clinical |
| Seven Bridges | Precision medicine | No | NCI Cancer Genomics Cloud | SOC 2, HIPAA | Enterprise licensing | Oncology teams |
| Code Ocean | Reproducibility | No | None (general-purpose) | Audit trails, provenance | Institutional licensing | Reproducibility and audits |
| Phylo.Bio | AI-native (orchestration) | Yes (conversational) | 68 via AI agents | TBD | Free preview / custom | Tool orchestration breadth |
| K-Dense | AI-native (autonomous) | Yes (multi-agent) | Literature-focused | TBD | $199-$499/mo + custom | Autonomous scientific research |
| Purna AI MIP | AI-native (biology) | Yes (natural language) | 30+ live databases | In development | Research credits available | Biology-specific reasoning |
How to Choose: A Decision Framework
Picking a platform starts with understanding what your team actually needs. Not what sounds impressive in a demo, but what solves the problems your researchers face daily.
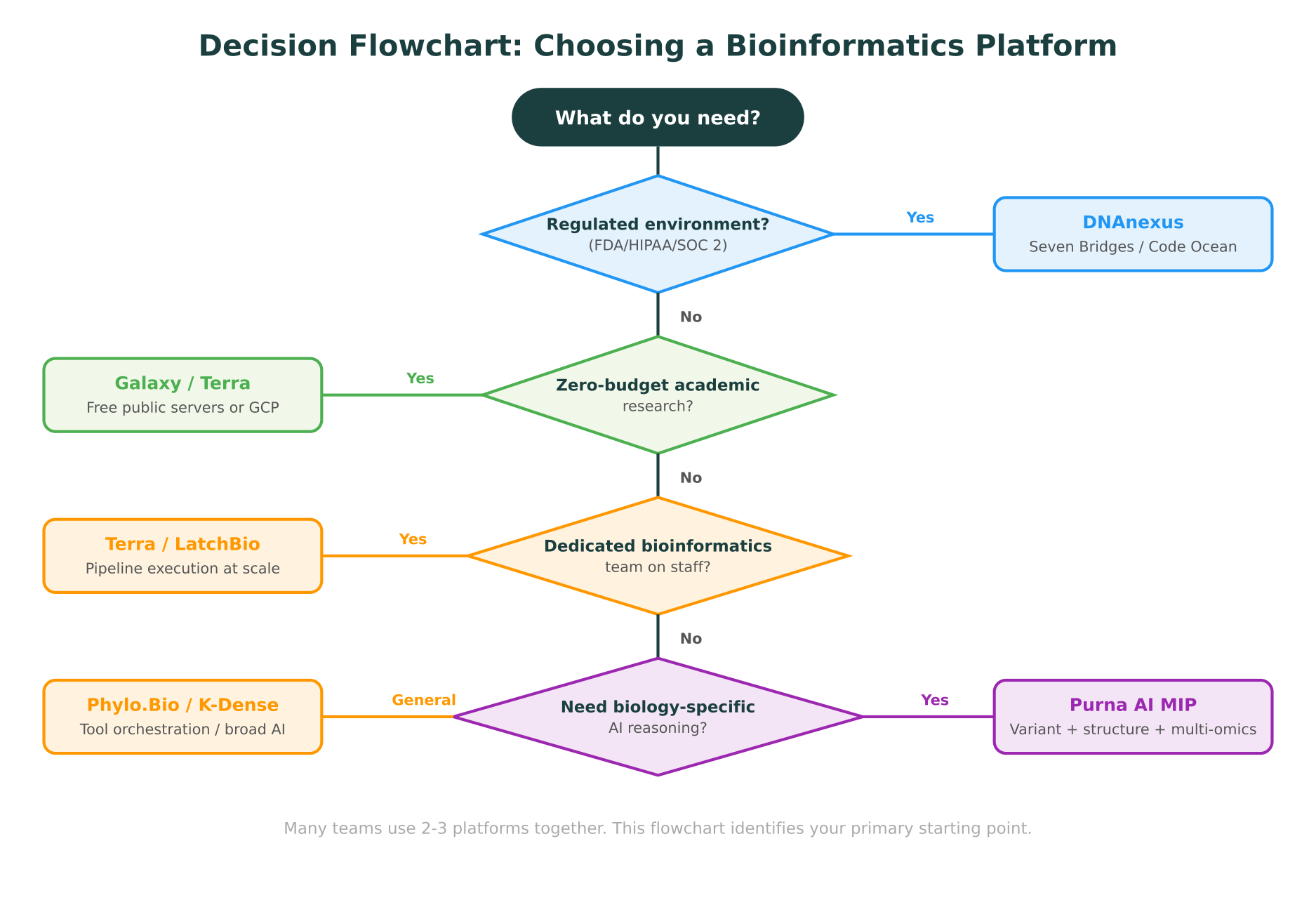
Start with compliance. If your organization processes clinical samples, operates under FDA oversight, or handles data subject to HIPAA or GDPR, your platform choice is constrained. DNAnexus and Seven Bridges have spent years building the certifications and validated environments these settings require. Choosing a platform without the right compliance posture is not a risk worth taking.
Evaluate your team’s technical depth. If you have a dedicated bioinformatics team comfortable with WDL, CWL, or Nextflow, platforms like Terra and Galaxy give you maximum control and flexibility. If your team is primarily wet-lab scientists who need results without managing pipelines, the AI-native platforms (Phylo.Bio, K-Dense, MIP) reduce the barrier significantly.
Consider your analysis types. Population-scale genomics with thousands of samples points toward Terra. Cancer genomics with diverse populations points toward Seven Bridges. Variant interpretation with structural analysis points toward MIP. Reproducibility for publications points toward Code Ocean. No single platform covers everything perfectly.
Think about the intelligence layer. This is the new dimension in platform selection. Traditional platforms run your pipelines and return results. AI-native platforms reason about your data, synthesize evidence from multiple sources, and provide interpretations with citations. If your researchers spend significant time manually searching ClinVar, cross-referencing gnomAD, and reading literature to interpret results, an AI-native platform can compress that workflow substantially.
Plan for integration, not replacement. Most serious bioinformatics operations will use two or three platforms. A common pattern emerging in 2026: Terra or Galaxy for pipeline execution, an AI-native platform like MIP for interpretation and reasoning, and Benchling for lab informatics. Choosing a primary platform does not mean choosing an only platform.
Closing Thoughts
The best bioinformatics platforms in 2026 span a wider range than ever before. From Galaxy’s community-driven open-source ecosystem to DNAnexus’s enterprise compliance infrastructure to the new AI-native platforms that reason about biology, the options reflect the growing diversity of what computational biology teams need.
The most significant shift is the emergence of platforms that provide intelligence, not just infrastructure. Running a pipeline is necessary but not sufficient. What matters increasingly is what happens after the pipeline finishes: interpreting variants, understanding structural impacts, connecting findings across databases, and reaching defensible biological conclusions. The platforms that help researchers move from raw data to insight, with evidence and citations at every step, will define the next era of computational biology.
For biology teams ready to explore what an AI-native, evidence-grounded workspace looks like in practice, visit Purna’s Molecular Intelligence Platform.
Purna’s Molecular Intelligence Platform provides AI-powered variant interpretation, 3D protein structure analysis, containerized code execution, and natural language access to 30+ biological databases. Every answer is cited to source data, not training data.
Explore Purna's Molecular Intelligence Platform
AI-powered workspace for biology teams to accelerate drug discovery from target identification to lead optimization.
Try Purna AI →
