Open-Source vs Commercial Bioinformatics Platforms: Which Is Right for Your Lab?

The question of whether to use open-source or commercial bioinformatics tools has no universal answer. It depends on your lab’s size, technical capacity, regulatory requirements, and what you actually need the software to do. The right choice for a computational genomics core facility with three bioinformaticians on staff is not the right choice for a clinical lab processing patient samples under CLIA/CAP oversight, and neither is the right choice for a five-person academic research group on a tight NIH budget.
This post provides an honest, structured comparison. We cover the major open-source platforms (Galaxy, Nextflow, Snakemake, Bioconductor), the established commercial options (Terra, DNAnexus, Seven Bridges), and the newer category of AI-powered platforms like Purna’s Molecular Intelligence Platform (MIP). For each, we examine total cost of ownership, compliance readiness, scalability, learning curves, and support models. At the end, we provide a decision framework that lab leads can use to evaluate which approach fits their specific situation.
The Open-Source Landscape: What You Get and What You Build
Open-source bioinformatics tools have been the backbone of computational biology for decades. They are powerful, flexible, and free to download. They are also, in many cases, far more expensive to operate than their sticker price suggests.
Galaxy: the web-based workbench
Galaxy, first released in 2005 and continuously maintained by a global community, provides a web-based graphical interface for running bioinformatics analyses. Users can chain tools into workflows, share analyses with collaborators, and access a library of over 9,000 tools covering genomics, proteomics, metabolomics, and more.
Strengths. Galaxy lowers the barrier to entry compared to pure command-line work. Its workflow system supports reproducibility. The community is active, with regular training events, a dedicated training infrastructure (Galaxy Training Network), and public servers that let researchers run analyses without local compute. For academic labs that need to run standard pipelines (RNA-seq, variant calling, ChIP-seq) without managing their own infrastructure, Galaxy’s public instances can be a genuine zero-cost starting point.
Limitations. Galaxy’s public servers have resource limits that make large-scale analyses impractical. Running your own Galaxy instance requires server administration, tool installation, dependency management, and ongoing maintenance. A 2023 community survey found that institutional Galaxy administrators spent an average of 15 to 20 hours per week on maintenance tasks. The web interface, while more accessible than a command line, still requires understanding of tool parameters, data formats, and analysis logic. It is not “no-code” in the way a bench scientist typically needs.
Nextflow and Snakemake: pipeline orchestration
Nextflow and Snakemake occupy a different niche. They are workflow management systems that let bioinformaticians define analysis pipelines as code, then execute them reproducibly across different compute environments (local machines, HPC clusters, cloud platforms).
Strengths. Both tools excel at reproducibility and portability. Nextflow’s nf-core repository contains over 90 peer-reviewed, community-maintained pipelines for common bioinformatics tasks. Snakemake integrates tightly with conda environments and container runtimes. For bioinformatics teams building custom pipelines, these tools represent best-in-class workflow management.
Limitations. Nextflow and Snakemake are tools for bioinformaticians, not for end-user researchers. Writing a Nextflow pipeline requires proficiency in Groovy (or at minimum, DSL2 syntax). Snakemake uses Python-like syntax but still demands programming skills. These tools solve the reproducibility and scalability problem for teams that already have the technical expertise. They do not solve the accessibility problem for researchers who lack coding skills.
Bioconductor: the R ecosystem
Bioconductor provides a curated collection of R packages for the analysis and comprehension of high-throughput genomic data. With over 2,200 software packages, it covers everything from differential expression analysis to single-cell RNA-seq to flow cytometry.
Strengths. Bioconductor’s statistical rigor is well-established. Packages like DESeq2, edgeR, and Seurat are standard tools in their respective fields. The vignette-based documentation is often excellent. For researchers who are comfortable in R, Bioconductor is an indispensable resource.
Limitations. Bioconductor requires R programming skills. Package management across R versions and operating systems can be fragile. Scaling beyond a single workstation typically requires additional infrastructure (HPC access, cloud configuration). And integrating Bioconductor analyses with other tools (variant annotation, protein structure analysis, database queries) requires custom scripting.
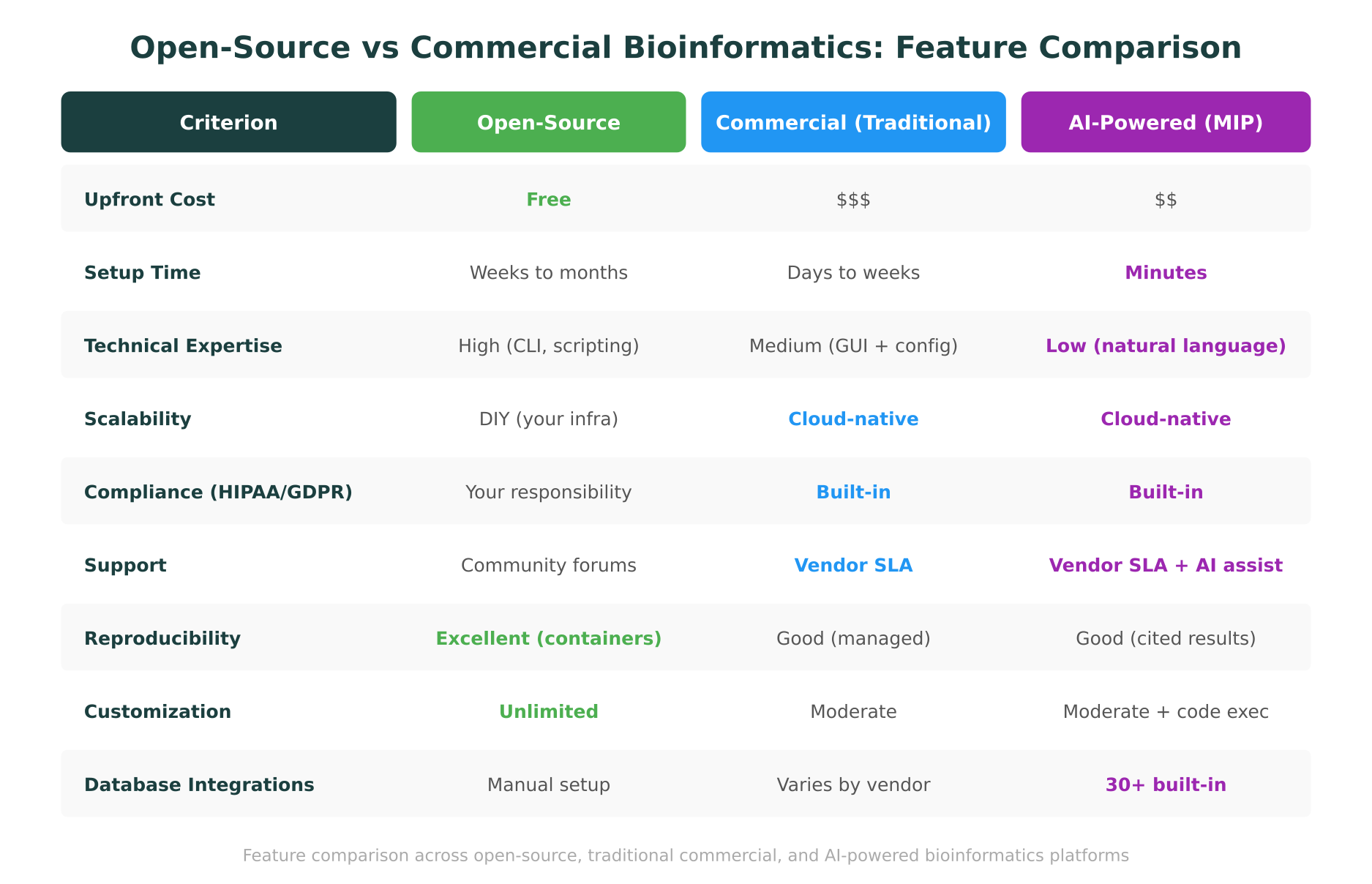
The Commercial Landscape: What You Buy and What You Avoid
Commercial bioinformatics platforms emerged to address the operational burden that open-source tools place on research teams. They trade software freedom for managed infrastructure, compliance guarantees, and vendor support.
Terra: cloud-native genomics at scale
Terra, developed by the Broad Institute and Verily, runs on Google Cloud Platform and provides a web-based environment for running genomic analyses. It supports WDL (Workflow Description Language) pipelines, Jupyter notebooks, and integrates with major genomic datasets (TCGA, gnomAD, the All of Us Research Program).
Strengths. Terra handles cloud infrastructure management, including compute provisioning, storage, and cost optimization. Its integration with the Broad’s GATK best practices pipelines makes it a strong choice for germline and somatic variant calling at scale. The data model supports large cohort studies. For institutions running thousands of samples through standardized pipelines, Terra provides real operational value.
Limitations. Terra’s learning curve is steeper than its marketing suggests. Configuring workspaces, writing or modifying WDL workflows, and managing cloud costs all require technical sophistication. Pricing can be unpredictable, as costs scale with compute usage on GCP. The platform is tightly coupled to Google Cloud, which can be a concern for institutions with multi-cloud or on-premise requirements. And while Terra excels at pipeline execution, it does not provide the kind of AI-powered analysis and interpretation that newer platforms offer.
DNAnexus: enterprise genomics
DNAnexus positions itself as an enterprise platform for genomic data management and analysis. It supports multiple cloud backends (AWS and Azure), provides FDA-compliant environments, and offers extensive APIs for integration with existing systems.
Strengths. DNAnexus has strong compliance credentials, with SOC 2, HIPAA, and FedRAMP certifications. The platform supports both graphical workflow building and command-line access. Its data management capabilities, including access controls, audit logging, and data provenance tracking, are well-suited to clinical and regulated environments.
Limitations. DNAnexus is priced for enterprise customers. Academic labs and smaller research groups often find the licensing costs prohibitive. The platform requires meaningful onboarding, and building custom analyses still demands programming skills (typically Python or R via the DNAnexus API or app framework). Support is responsive but tied to contract tiers.
Seven Bridges: workflow execution and data management
Seven Bridges provides a cloud platform for bioinformatics analysis, with particular strength in workflow execution using CWL (Common Workflow Language). It also operates the Cancer Genomics Cloud, one of the NCI’s cloud resources for accessing and analyzing cancer genomic data.
Strengths. Seven Bridges’ CWL support and integration with NCI datasets make it valuable for cancer genomics research. The platform offers compliance certifications and managed infrastructure.
Limitations. Like Terra and DNAnexus, Seven Bridges requires technical proficiency to use effectively. The CWL learning curve is comparable to WDL. Pricing is opaque and generally enterprise-oriented.
The Hidden Costs of “Free” Software
The most common mistake in open-source vs commercial bioinformatics comparisons is equating “free software” with “no cost.” Open-source tools have zero licensing fees. They do not have zero cost.
Infrastructure
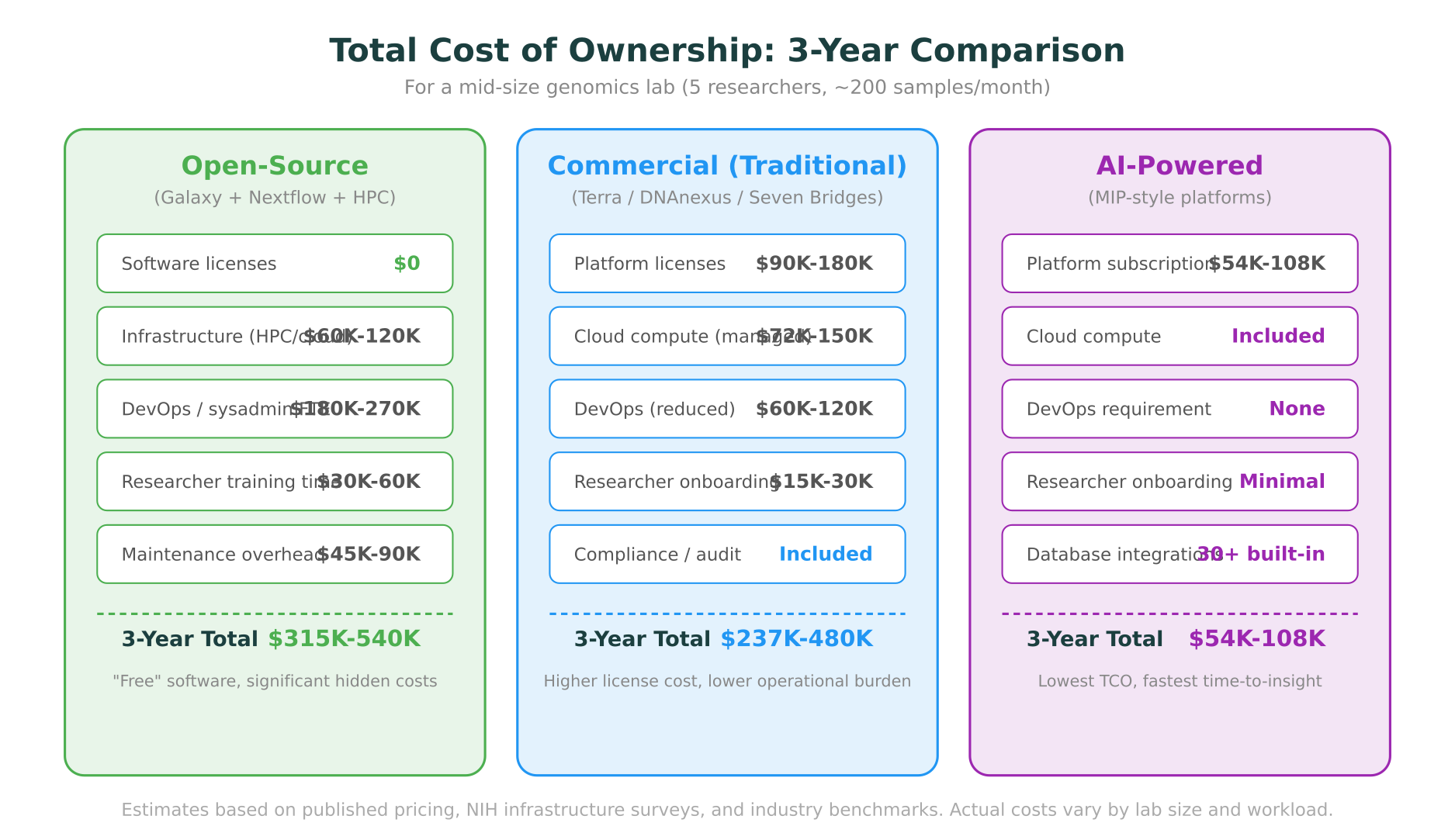
Running bioinformatics workloads requires compute and storage. For open-source tools, this means either maintaining on-premise HPC infrastructure or configuring cloud resources. A mid-size genomics lab processing 200 samples per month will spend $20,000 to $40,000 per year on cloud compute alone, before accounting for storage, data transfer, and the human time to manage it.
Personnel
Open-source bioinformatics infrastructure needs people to maintain it. Installing and updating tools, managing dependencies, configuring compute resources, troubleshooting failures, and training users all require staff time. A 2024 survey by the NIH Office of Data Science Strategy found that 62% of institutional bioinformatics cores employed at least one full-time systems administrator dedicated to maintaining analysis infrastructure.
For labs without dedicated IT support, this burden falls on the bioinformaticians themselves, diverting time from research to system administration.
Training
Every open-source tool has a learning curve. Galaxy is simpler than the command line, but researchers still need training. Nextflow and Snakemake require programming skills. Bioconductor requires R proficiency. The time researchers spend learning tools is time not spent on research. For a PI evaluating total cost of ownership, researcher training hours have a real dollar value.
Opportunity cost
Perhaps the most significant hidden cost is what does not get done. When a research group lacks the technical capacity to run analyses, hypotheses go untested. When clinical labs cannot scale variant interpretation, turnaround times increase. When a structural biologist needs to understand a protein mutation but cannot run the computational analysis, they wait for a collaborator or skip the analysis entirely.
These opportunity costs do not appear on any invoice, but they are real.

Where AI-Powered Platforms Fit In
A newer category of bioinformatics platform has emerged that addresses some limitations of both open-source and traditional commercial options. These platforms use AI not just as a feature but as the primary interface for interacting with biological data and analyses.
Purna AI’s Molecular Intelligence Platform (MIP) is one example. Rather than providing a pipeline runner or a workflow builder, MIP provides a natural language interface where researchers describe what they need, and the platform handles database queries, code execution, and evidence synthesis.
What this changes
For labs without bioinformatics staff. A wet lab researcher can ask MIP to classify a variant, analyze a protein structure, or query clinical databases without writing code or configuring tools. The platform connects to over 30 databases (ClinVar, gnomAD, OMIM, UniProt, PDB, and others), runs analyses in containerized environments, and returns cited results. This addresses the accessibility gap that open-source tools have not solved, as discussed in our post on bioinformatics without coding.
For labs with bioinformatics staff. MIP does not replace technical expertise. It augments it. Bioinformaticians can use the code execution environment for custom analyses, inspect and modify generated code, and use the natural language interface for rapid exploration across databases. The platform handles the routine work (database queries, standard analyses, evidence compilation) so that skilled staff can focus on novel computational problems.
For clinical labs. Variant interpretation is one of MIP’s core capabilities. The platform applies ACMG/AMP criteria with structured reasoning, pulling evidence from multiple databases and providing citations for every classification. This does not replace expert review, but it accelerates the initial classification and ensures no relevant data source is missed.
What this does not change
AI-powered platforms are not the right choice for every use case. Labs that need to run custom Nextflow pipelines on thousands of whole genomes are better served by purpose-built pipeline infrastructure. Research groups developing novel bioinformatics methods need the full flexibility of open-source environments. And no AI platform should be trusted to make clinical decisions without human expert oversight.
The value proposition of AI-powered platforms is clearest for teams that need to do analysis and interpretation across multiple omics domains but lack the technical staff to build and maintain the infrastructure themselves. For these teams, the question is not “open-source vs commercial” but “how much of our budget should go to infrastructure vs. actual research?”
Compliance and Regulatory Considerations
For clinical and translational labs, compliance is often the deciding factor in platform selection.
Open-source: your responsibility
Using open-source tools in a regulated environment is possible but demanding. HIPAA compliance requires encryption at rest and in transit, access controls, audit logging, and breach notification procedures. GDPR adds data subject rights and cross-border transfer restrictions. Building these capabilities on top of open-source tools requires security expertise, documented procedures, and regular audits.
Some institutions solve this by running open-source tools within a compliant infrastructure layer (e.g., a HIPAA-compliant AWS or GCP environment), but the configuration and ongoing compliance monitoring remain the lab’s responsibility.
Commercial: largely handled
Established commercial platforms like Terra, DNAnexus, and Seven Bridges invest heavily in compliance certifications. SOC 2, HIPAA BAAs, FedRAMP, and in some cases ISO 27001 certifications transfer much of the compliance burden from the lab to the vendor. This is a significant value proposition for clinical labs where the cost of a compliance failure far exceeds any software licensing fee.
AI-powered platforms: evolving
Newer platforms are actively building compliance capabilities. For any lab handling patient data, the critical questions are: where is data stored, who has access, what certifications does the vendor hold, and how are AI-generated results documented for audit trails?
Scalability: From 10 Samples to 10,000
Scale requirements vary enormously across labs, and they significantly influence platform choice.
Open-source tools scale horizontally but require engineering. Nextflow and Snakemake can distribute work across cloud instances or HPC nodes, but someone needs to configure the compute backend, manage resource allocation, and handle failures. Galaxy’s public servers have hard resource limits.
Commercial platforms provide managed scaling. Terra, DNAnexus, and Seven Bridges handle autoscaling, resource provisioning, and cost optimization as core features. For large-scale genomics projects (population studies, clinical sequencing programs), this managed scaling justifies the platform cost.
AI-powered platforms are designed for different scale patterns. Rather than processing thousands of samples through a single pipeline, platforms like MIP handle the complexity of querying multiple databases and synthesizing evidence across data types. The scale challenge they solve is analytical breadth (pulling from 30+ databases per query) rather than sample throughput.
A Decision Framework for Lab Leads
Choosing a bioinformatics platform should start with an honest assessment of your lab’s needs and capabilities. Here is a structured approach.
Step 1: Assess your technical capacity
- Do you have dedicated bioinformatics staff? (Not “a postdoc who knows some Python,” but someone whose primary role is computational.)
- Can those staff maintain infrastructure, or do they spend most of their time on analysis?
- How technically skilled are your end-user researchers?
If you have strong bioinformatics staff with infrastructure skills, open-source tools can be highly cost-effective. If your bioinformaticians should be doing analysis rather than system administration, a managed platform (commercial or AI-powered) frees them for higher-value work. If your researchers are primarily bench scientists, an AI-powered platform with a natural language interface may be the only option that gets adopted.
Step 2: Define your regulatory requirements
- Do you handle patient data? (HIPAA, GDPR, local equivalents)
- Do you need audit trails for clinical reporting?
- Are you subject to CLIA/CAP or equivalent laboratory standards?
If compliance is a hard requirement, the cost of building it on open-source infrastructure is substantial. Commercial and AI-powered platforms with existing certifications are usually more practical.
Step 3: Estimate your actual workload
- How many samples per month?
- How many distinct analysis types?
- Do you need primarily pipeline execution, or analysis and interpretation?
High-throughput pipeline execution favors Nextflow/Snakemake (with appropriate infrastructure) or commercial pipeline platforms. Analysis, interpretation, and database queries favor AI-powered platforms. Many labs need both.
Step 4: Calculate total cost of ownership
Include not just software and compute costs, but also:
- Staff time for infrastructure maintenance
- Researcher time for training and tool learning
- Opportunity cost of analyses not performed
- Compliance implementation and ongoing audit costs
A platform that costs $1,500/month but saves a bioinformatician 20 hours/week of infrastructure work is a net gain for most labs. A “free” tool that requires a dedicated system administrator at $90K/year is not actually free.
Step 5: Consider the hybrid approach
Many successful labs use a combination. A common pattern: Nextflow for high-throughput pipeline execution, plus an AI-powered platform like MIP for variant interpretation, database queries, and exploratory analysis. This gives you the pipeline flexibility of open-source tools with the analytical depth and accessibility of a managed platform.
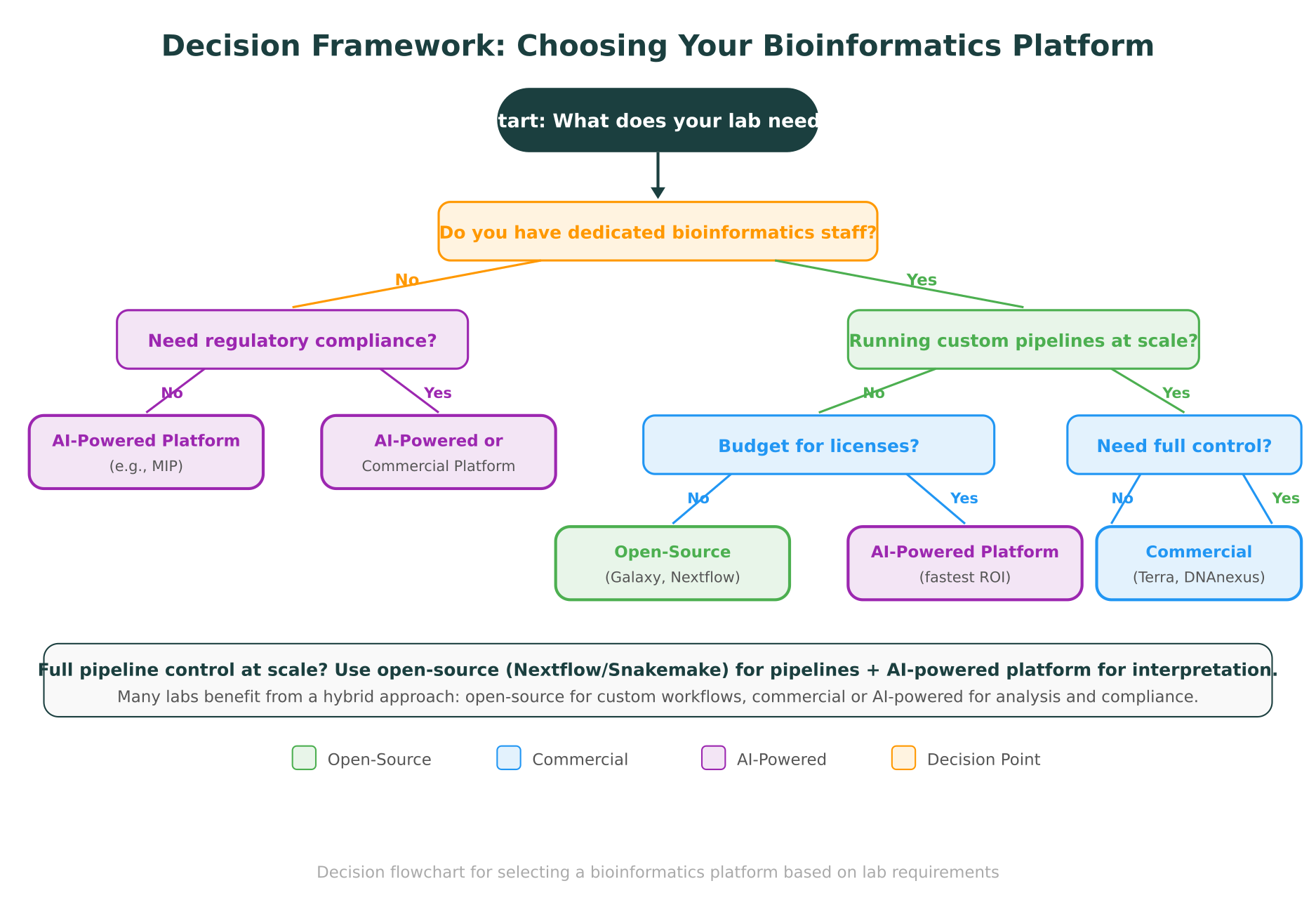
What the Landscape Looks Like in 2026
The bioinformatics platform landscape is converging. Open-source tools are adding cloud-native features (Nextflow Tower, Galaxy on Kubernetes). Commercial platforms are incorporating AI capabilities. AI-powered platforms are expanding their pipeline execution features.
The most significant shift is the emergence of molecular intelligence as a concept: platforms that do not just run computations but reason across data types, databases, and analytical frameworks. This is different from adding a chatbot to a pipeline runner. It represents a fundamentally different architecture where AI is the connective layer between data sources, analytical tools, and domain knowledge.
For lab leads making platform decisions today, the practical implication is that the “build vs. buy” calculus has changed. Five years ago, building on open-source was almost always the right call for labs with technical staff. Today, the complexity of integrating across 30+ databases, maintaining compliance, and providing accessible interfaces for diverse research teams makes the case for managed platforms stronger than it has ever been.
This does not mean open-source tools are obsolete. For custom pipeline development, novel method implementation, and situations where full control over every parameter is essential, open-source remains the right choice. But for the growing majority of labs that need to perform multi-database analyses, interpret variants, analyze protein structures, and do it all while meeting compliance requirements, the total cost of doing it yourself increasingly exceeds the cost of using a platform designed for the purpose.
Making the Decision
The right platform depends on your lab, not on ideology. Open-source tools are not inherently better because they are free, and commercial tools are not inherently better because they cost money. What matters is whether the platform matches your team’s skills, your regulatory obligations, your scale requirements, and your budget when all costs are counted.
Start with the decision framework above. Be honest about your team’s technical capacity. Count the hidden costs. And remember that the best bioinformatics platform is the one your researchers actually use, not the one that looks best in a grant application.
For teams exploring AI-powered bioinformatics for the first time, Purna AI offers up to $10,000 in research credits to qualified research groups. It is a low-risk way to evaluate whether a molecular intelligence approach fits your lab’s workflow before committing to a platform decision.
Purna AI’s Molecular Intelligence Platform integrates 30+ clinical and biological databases, AI-powered variant interpretation, 3D protein structure analysis, and containerized code execution in a single workspace. Researchers can query databases, run analyses, and interpret results through natural language, with every conclusion cited to its source. Apply for research credits to try it with your own data.
Explore Purna's Molecular Intelligence Platform
AI-powered workspace for biology teams to accelerate drug discovery from target identification to lead optimization.
Try Purna AI →
