AI for Rare Disease Diagnosis: Current Progress and What Comes Next

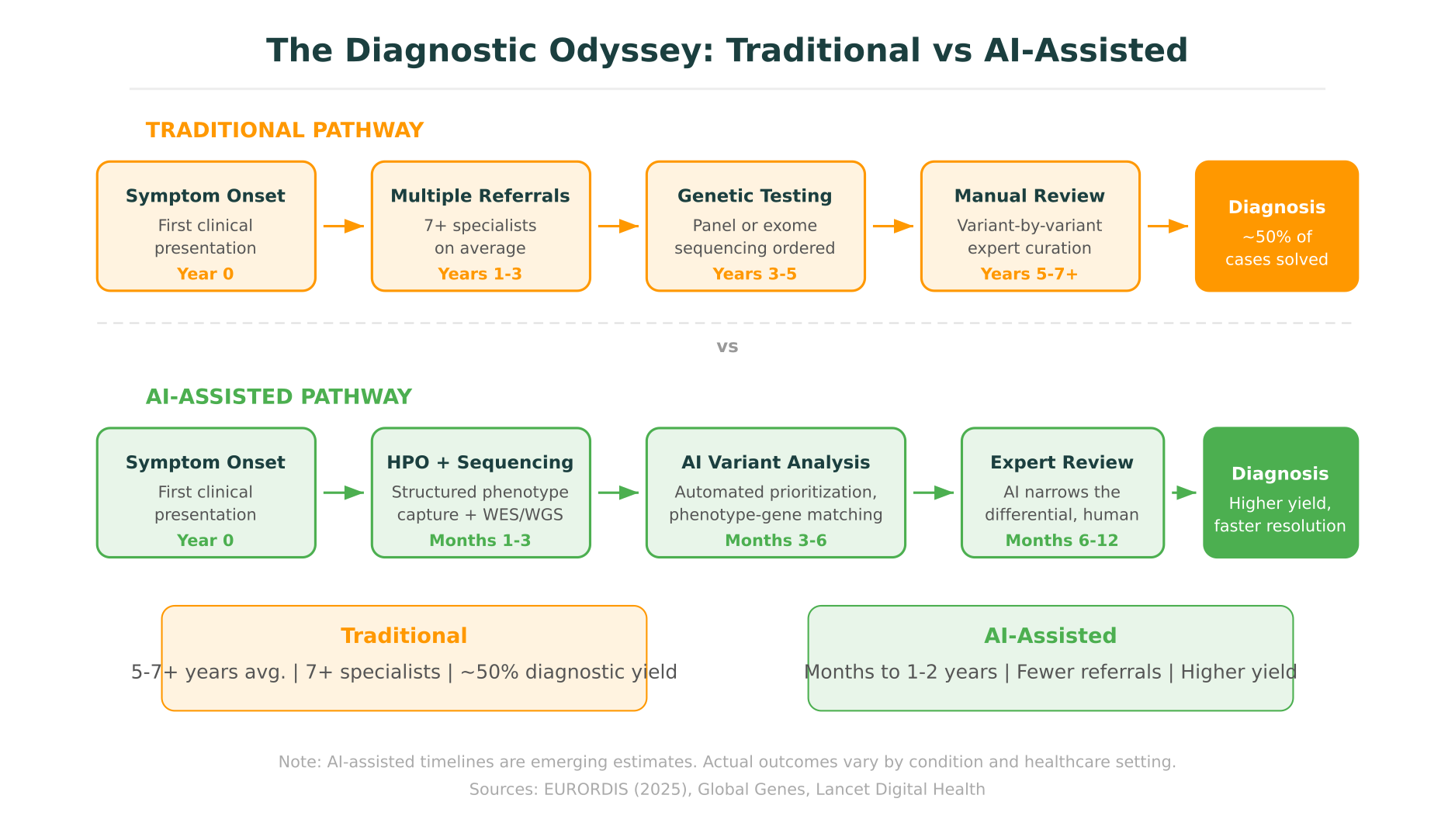
For the roughly 300 million people worldwide living with a rare disease, getting a diagnosis is often the hardest part. The average time from first symptoms to a confirmed diagnosis exceeds five years. Patients see seven or more specialists along the way. And approximately 30% never receive a definitive diagnosis at all. This is what clinicians and patient advocacy groups call the “diagnostic odyssey,” a problem that has persisted for decades despite advances in genomic sequencing.
AI rare disease diagnosis is now emerging as a serious area of progress. Not a solved problem, not a replacement for clinical expertise, but a set of tools that are measurably shortening the time to diagnosis and increasing diagnostic yield in specific contexts. The most notable development in 2026 is DeepRare, an agentic AI system published in Nature that outperformed experienced clinicians on diagnostic benchmarks while also revealing just how far the field still has to go.
This post covers what is working, what is not, and where AI for rare diseases is realistically heading.
The Diagnostic Odyssey: Why Rare Diseases Are So Hard to Diagnose
A disease is considered “rare” when it affects fewer than 200,000 people in the United States (or fewer than 1 in 2,000 in the EU). Individually, each condition is uncommon. Collectively, the numbers are staggering: over 7,000 known rare diseases affect an estimated 300 million people globally, according to Global Genes and EURORDIS. Roughly 72% of rare diseases are genetic in origin, and about 70% manifest in childhood.
The diagnostic challenge comes from several converging factors.
Limited clinical familiarity
Most physicians encounter any given rare disease infrequently, if ever. Medical education covers common conditions in depth but can only scratch the surface of the thousands of rare syndromes. A pediatric neurologist might see one case of a specific lysosomal storage disorder across an entire career.
Phenotypic overlap and heterogeneity
Many rare diseases share symptoms with common conditions, leading to initial misdiagnoses. Conversely, the same genetic condition can present very differently across patients, even within the same family. A single-gene disorder might cause intellectual disability in one patient and seizures in another, making pattern recognition difficult even for specialists.
Variant interpretation bottleneck
Even when whole-exome sequencing (WES) or whole-genome sequencing (WGS) is ordered, the interpretation step remains a bottleneck. A typical exome produces roughly 20,000 to 30,000 variants. Filtering these down to the one or two that explain a patient’s condition requires cross-referencing population frequency databases, clinical variant databases, protein function predictors, literature reports, and phenotype ontologies. This process is labor-intensive, and the volume of variants of uncertain significance (VUS) in rare disease contexts is particularly high because many pathogenic variants in rare conditions have never been reported before.
DeepRare: What an Agentic AI System Can and Cannot Do
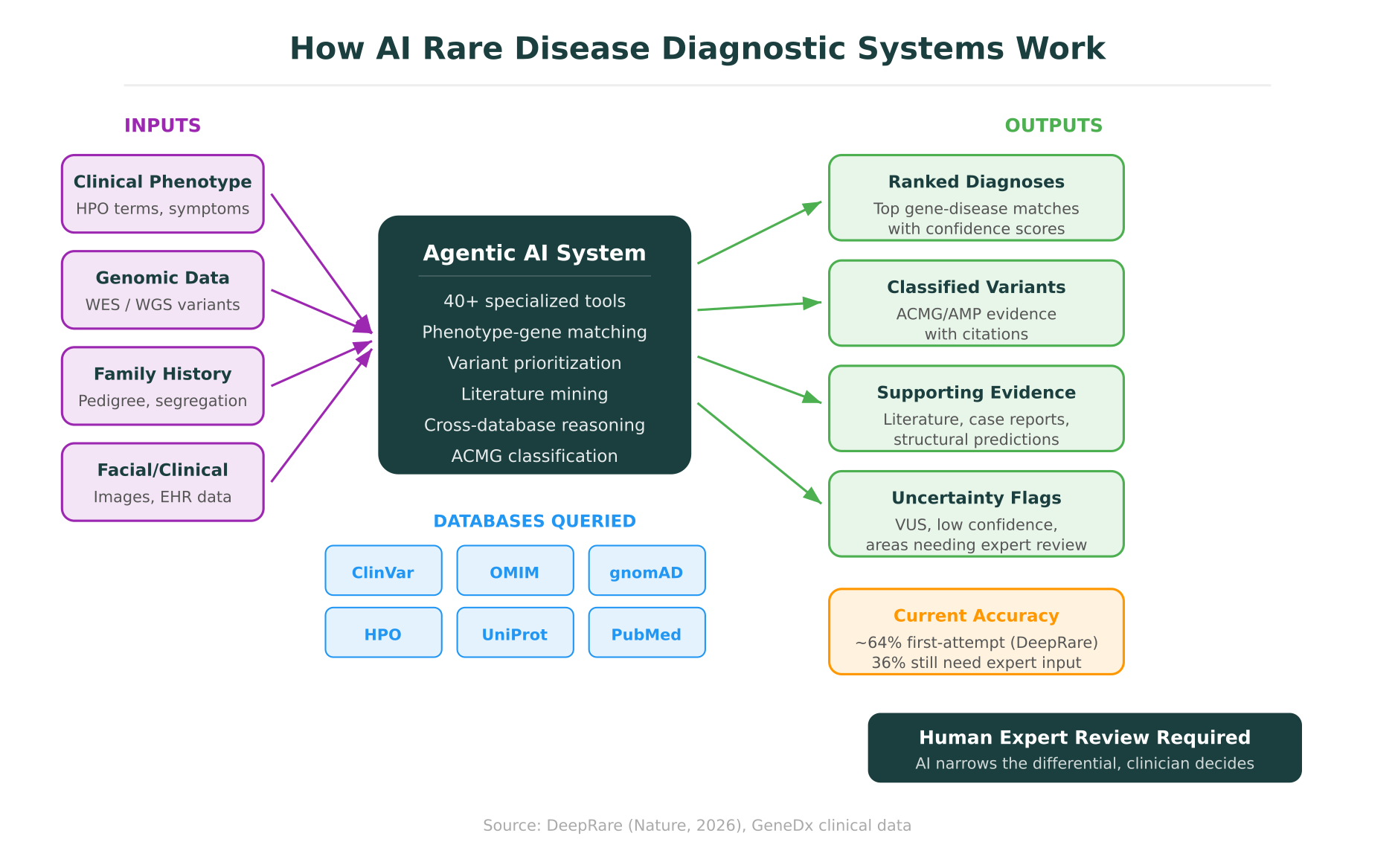
The most significant development in AI rare disease diagnosis this year is DeepRare, published in Nature in early 2026. Unlike earlier tools that tackled isolated steps of the diagnostic process, DeepRare is an agentic system: it orchestrates 40+ specialized tools to work through the full diagnostic workflow, from phenotype encoding to variant prioritization to gene-disease matching.
How DeepRare works
DeepRare takes a patient’s clinical description (symptoms, family history, lab findings) and genomic data as input. It then autonomously decides which tools to invoke and in what sequence. These tools include phenotype encoders that map clinical descriptions to Human Phenotype Ontology (HPO) terms, variant annotation engines, population frequency databases, gene-disease association databases (including OMIM and ClinGen), protein structure predictors, and literature search tools.
The system does not simply look up answers. It reasons iteratively: if an initial set of candidate genes does not adequately explain the phenotype, it reranks, queries additional databases, and considers alternative genetic mechanisms (compound heterozygosity, X-linked inheritance, de novo variants). This agentic loop, where the system plans, executes, evaluates, and replans, is what distinguishes DeepRare from earlier pipeline-based approaches.
Benchmark results: impressive but incomplete
On standardized rare disease diagnostic benchmarks, DeepRare outperformed teams of experienced clinical geneticists. Its first-attempt diagnostic accuracy was approximately 64%, and when allowed to refine its answers iteratively, it solved additional cases that clinicians missed on initial review.
That 64% figure deserves careful examination. It means that on the first pass, roughly 36% of cases were not correctly diagnosed. For conditions where the phenotype maps cleanly to a well-characterized Mendelian gene, the system performs well. For cases involving novel gene-disease associations, complex multigenic contributions, or extremely rare conditions with limited published case reports, accuracy drops considerably.
What the 36% tells us
The cases DeepRare struggles with are often the same ones that stump human experts: patients with atypical presentations, conditions caused by variants in genes not yet associated with disease, cases where the pathogenic variant is in a non-coding region not covered by exome sequencing, and situations where the phenotype results from the interaction of multiple genetic and environmental factors.
This is not a criticism of the system. It is a reflection of where the field stands. Even the best human diagnostic teams do not achieve 100% diagnostic yield. The current benchmark for experienced clinical geneticists reviewing rare disease cases is roughly 25-40% diagnostic yield from initial exome sequencing, depending on the patient population and the rigor of phenotypic data collection. DeepRare’s 64% represents a meaningful improvement over this baseline, but it also makes clear that AI is not yet solving the hardest cases on its own.
GeneDx and Large-Scale Rare Disease Genomics
While DeepRare demonstrates what is possible in a research setting, GeneDx shows what AI-assisted rare disease diagnosis looks like at clinical scale.
GeneDx is one of the largest clinical genomics laboratories focused on rare disease, with a dataset of nearly one million exomes and genomes, approximately one million uniquely classified variants, and over seven million phenotypic data points collected over more than a decade of clinical testing.
AI-powered variant ranking
GeneDx has integrated AI-based variant ranking into its clinical workflow. Their system uses machine learning models trained on their proprietary dataset to prioritize variants most likely to be pathogenic for a given patient’s phenotype. This does not replace the human review step, where board-certified geneticists and genetic counselors evaluate the top candidates, but it substantially reduces the time and cognitive load of the initial triage.
Their reported diagnostic yield of 51.4% across more than 300,000 cases is notable for a real-world clinical dataset, significantly higher than the 25-35% yield typically reported for exome sequencing in the literature. Part of this is attributable to their variant ranking AI; part reflects the advantage of an internal database that grows with every case processed.
The data flywheel
GeneDx illustrates a pattern that is becoming central to AI in rare disease: the data flywheel. Every case that goes through their system, whether it results in a diagnosis or not, adds to the training data. Variants classified in one patient can inform classification in the next. Phenotypic patterns observed across thousands of patients with the same condition sharpen the model’s ability to recognize atypical presentations.
This creates an advantage that is difficult for smaller labs or academic centers to replicate. The more cases a system has seen, the better it becomes at recognizing rare variants and atypical presentations. This is one reason why the landscape of AI-assisted rare disease diagnosis is likely to consolidate around a relatively small number of large-scale platforms and databases, a dynamic explored in more detail in our post on open-source vs. commercial bioinformatics platforms.
How AI Helps in Rare Disease Diagnosis
AI tools for rare disease do not follow a single approach. Several distinct methods have shown clinical utility, each addressing a different part of the diagnostic bottleneck.
Phenotype-to-gene matching
The Human Phenotype Ontology (HPO) provides a standardized vocabulary for describing clinical features. Tools like Exomiser, PhenoDB, and LIRICAL use HPO terms to compute similarity scores between a patient’s phenotype and the known phenotypic profiles of thousands of genetic conditions. When a clinician enters a structured phenotype (e.g., “intellectual disability + seizures + microcephaly + hypotonia”), these tools generate a ranked list of candidate genes.
DeepRare improved on this approach by using a language model to extract HPO terms from unstructured clinical notes, reducing the manual coding burden that has historically limited the adoption of phenotype-driven tools.
Variant prioritization in rare disease contexts
Standard variant filtering pipelines work well for common disease-associated variants but are less effective in rare disease, where the causal variant may be novel (never previously reported), in a gene with limited functional characterization, or present at low frequency even in large population databases.
AI-based variant prioritization for rare disease goes beyond simple frequency filtering. It incorporates protein structure predictions, evolutionary conservation scores, splice site impact models, and gene-level constraint metrics to assess whether a novel variant in a poorly characterized gene is likely to be damaging. This is the kind of integrative analysis that benefits from having multiple data sources available in a single workspace, the same principle behind molecular intelligence platforms that connect variant databases, structural models, and functional predictors.
Facial gestalt analysis
Several rare genetic syndromes produce characteristic facial features, subtle enough that many clinicians would not recognize them but consistent enough for pattern recognition. Face2Gene, developed by FDNA, and the more recent GestaltMatcher use deep learning models trained on facial photographs to suggest candidate syndromes.
GestaltMatcher, published in Nature Genetics, extends this approach to ultra-rare conditions by using a similarity-based matching framework rather than requiring hundreds of training examples for each syndrome. In validation studies, it correctly identified the syndrome for patients with conditions represented by as few as one training image.
These tools are not standalone diagnostic instruments. They generate candidate lists that inform the clinical differential, which must then be confirmed through genetic testing and expert evaluation.
Literature mining for case reports
For ultra-rare conditions, the critical diagnostic clue may exist in a single published case report among millions of biomedical papers. AI-powered literature mining tools can search PubMed and preprint servers for phenotypic descriptions that match a patient’s presentation, surfacing relevant case reports that a manual search would likely miss.
This capability is particularly valuable when a patient has a variant in a gene that is not yet in OMIM or ClinGen as a disease-associated gene, but where one or two published case reports describe patients with similar variants and overlapping phenotypes. Platforms that integrate natural language querying of biological databases are making this kind of literature search more accessible to clinicians who do not have time to construct complex Boolean search queries.
Multi-omics integration
Genomic sequencing alone solves roughly half of rare disease cases. For the other half, additional data layers can provide the missing evidence. Transcriptomics (RNA sequencing) can reveal whether a suspected pathogenic variant actually affects gene expression or splicing. Proteomics can show whether the protein product is absent or dysfunctional. Metabolomics can identify biomarker patterns associated with specific inborn errors of metabolism.
AI models that integrate across these data types are still in early stages, but initial results are promising. In a study from the Broad Institute, RNA sequencing added diagnostic information in roughly 35% of cases that were negative on exome sequencing alone. The challenge, as with any multi-omics approach, is making the data integration tractable without requiring each patient to go through a research-grade analysis pipeline.
Current Limitations: An Honest Assessment
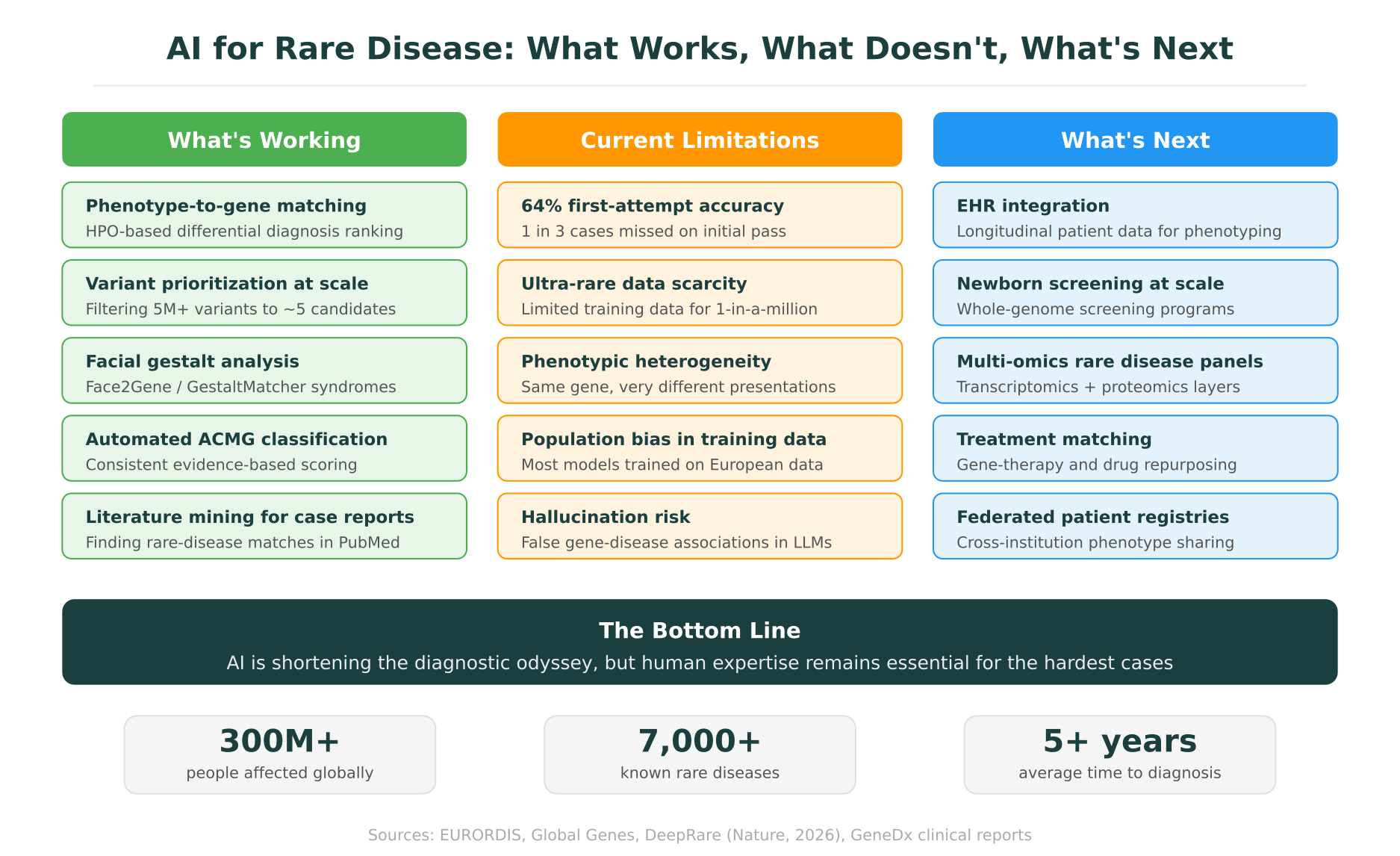
The progress is real, but the limitations are equally real. Responsible deployment of AI rare disease diagnosis tools requires acknowledging where these systems fall short.
Data scarcity for the rarest conditions
AI models learn from data, and for the rarest conditions, the data barely exists. There are over 7,000 known rare diseases, but hundreds of them have fewer than 50 documented patients in the published literature. For these ultra-rare conditions, there may not be enough examples for any AI model to learn the phenotype-genotype correlation reliably. A condition affecting 10 known patients worldwide cannot be learned from training data in the same way that a condition affecting 10,000 patients can.
Phenotypic heterogeneity
Even for well-studied rare diseases, the range of possible clinical presentations can be enormous. Marfan syndrome, caused by variants in the FBN1 gene, can manifest as anything from severe neonatal multi-organ disease to tall stature with mild joint hypermobility discovered incidentally in adulthood. AI models trained primarily on classic or severe presentations may miss patients with atypical or mild forms.
Population diversity gaps
The majority of genomic reference databases, training datasets, and published case reports are derived from individuals of European descent. This creates systematic blind spots. Variants that are common in African, South Asian, or East Asian populations may be poorly represented in population frequency databases, leading to incorrect prioritization. Phenotypic reference data may not capture how certain conditions present differently across ethnic groups.
GeneDx has noted this challenge in their own data, where diagnostic yield varies across populations, partly reflecting the uneven representation in reference databases. Addressing this requires not just algorithmic improvements but active expansion of genomic databases to include underrepresented populations.
Hallucination risk in gene-disease associations
Large language models (LLMs) used in agentic diagnostic systems can generate plausible-sounding but incorrect gene-disease associations. A system might confidently assert a link between a gene and a phenotype that does not exist in the curated literature, drawing instead from patterns in its training data that do not reflect validated biology.
DeepRare mitigates this by grounding its outputs in structured databases (OMIM, ClinGen, ClinVar) rather than relying solely on LLM-generated knowledge. But the risk is not eliminated. Any AI system that generates diagnostic hypotheses must be designed to surface its evidence sources and flag uncertainty, allowing the reviewing clinician to verify the reasoning. This is a principle that applies broadly across AI in clinical genomics, as discussed in our guide to reading genomic reports.
Regulatory uncertainty
Clinical diagnostic tools in the United States require FDA clearance or approval if they are marketed as diagnostic devices. The regulatory pathway for AI-based diagnostic systems in rare disease is still being defined. Most current tools are positioned as decision support rather than autonomous diagnostic instruments, which places them in a different regulatory category. But as these systems become more capable and more integrated into clinical workflows, the regulatory questions will become more pressing.
Where AI Rare Disease Diagnosis Is Heading
Several developments in the next two to five years are likely to reshape the landscape significantly.
EHR integration and longitudinal phenotyping
Most AI diagnostic tools today work from a static snapshot: a set of symptoms and a sequencing result at a single point in time. Integrating electronic health record (EHR) data would allow AI systems to track the evolution of a patient’s symptoms over months or years, capturing the progressive nature of many rare diseases. A patient who presents with developmental delay at age two and develops seizures at age five has a different diagnostic profile than one viewed at either time point alone.
Several health systems and EHR vendors are now piloting integrations that feed structured clinical data into rare disease diagnostic tools. The challenge is data standardization: EHR data is notoriously messy, and extracting reliable phenotypic information from clinical notes requires robust natural language processing.
Newborn screening with whole-genome sequencing
Multiple national programs are now piloting whole-genome sequencing as part of newborn screening. The UK’s Newborn Genomes Programme, BabySeq in the United States, and similar initiatives in Australia and the Middle East are generating the first large-scale data on what it looks like to screen newborns genomically for hundreds of rare conditions simultaneously.
AI is essential to making this feasible. No human team can manually interpret whole-genome sequencing results for every newborn. Automated variant filtering, phenotype-independent disease risk scoring, and automated ACMG classification are all necessary components. The ethical and clinical governance questions, particularly around incidental findings, carrier status, and adult-onset conditions, are still being worked through.
Treatment matching and drug repurposing
Diagnosis is not the end of the journey. For many rare diseases, even after a genetic cause is identified, no specific treatment exists. AI is beginning to contribute here as well, through drug repurposing models that identify existing approved drugs that target the same molecular pathway affected by a patient’s genetic variant.
This connects directly to the structural biology side of rare disease. Understanding how a specific variant alters protein structure and function can inform whether an existing drug that binds to or modulates that protein might be therapeutic. Platforms that integrate protein mutation impact prediction with variant classification and drug-target databases are well-positioned to support this kind of analysis.
Federated patient registries and data sharing
One of the most promising developments is the growth of federated patient registries that allow institutions to share phenotypic and genomic data without centralizing sensitive patient information. Networks like Matchmaker Exchange already connect rare disease databases across institutions, enabling clinicians to find patients with similar phenotypes and genotypes worldwide.
AI models trained across federated datasets can learn from a much larger patient population without any single institution needing to share raw data. This addresses both the data scarcity problem and the privacy concerns that have historically limited data sharing in clinical genomics.
Where Molecular Intelligence Platforms Fit
The rare disease diagnostic workflow touches nearly every capability that a comprehensive biology workspace needs: variant interpretation with ACMG classification, cross-referencing of clinical databases (ClinVar, OMIM, gnomAD, LOVD), structural analysis to assess the impact of novel missense variants, literature search for rare case reports, and phenotype-driven gene prioritization.
For many research and clinical teams, this means stitching together a dozen or more disconnected tools, an approach that is time-consuming and introduces opportunities for errors at every handoff. This is precisely the problem that molecular intelligence is designed to address: an integrated workspace where evidence from 30+ databases, structural predictions, and AI-powered interpretation come together in a single environment.
In a rare disease context, a clinician working in Purna’s MIP could query a novel variant and receive ACMG evidence criteria assembled from ClinVar, gnomAD population frequencies, protein domain annotations from UniProt, structural stability predictions from DynaMut2, and relevant case reports from the literature, all without switching tools or manually cross-referencing spreadsheets. For variants of uncertain significance, which are disproportionately common in rare disease cases, the ability to assess structural impact and functional consequences within the same workspace can provide the additional evidence needed to move toward a classification.
The platform does not replace expert judgment. A molecular intelligence workspace accelerates the evidence-gathering phase so that the clinician’s time is spent on the interpretive decisions that require human expertise rather than on the mechanical work of looking up database entries.
Conclusion
AI rare disease diagnosis has moved from a theoretical concept to a field with published benchmarks, clinical-scale implementations, and measurable improvements in diagnostic yield. DeepRare’s 64% first-attempt accuracy and GeneDx’s 51.4% clinical diagnostic yield represent genuine progress. They also represent how much work remains.
The diagnostic odyssey is getting shorter, not because AI has replaced the clinical geneticist, but because it is handling the parts of the workflow that do not require human judgment: encoding phenotypes, filtering variants, cross-referencing databases, mining literature. The hard cases, the ones involving novel gene-disease associations, atypical presentations, and ultra-rare conditions, still need experienced clinicians.
The most productive path forward is not AI alone or human expertise alone but a well-designed collaboration between the two. Systems that surface their evidence, flag their uncertainties, and leave the final diagnostic decision to the clinician are the ones most likely to earn trust and improve outcomes. For the 300 million people on a diagnostic odyssey, that collaboration cannot arrive soon enough.
MIP is Purna AI’s Molecular Intelligence Platform, an AI-powered workspace for biology teams. Variant interpretation, protein structure prediction, code execution in containerized environments, and 30+ database integrations in one place. Explore the platform at purna.ai. Eligible researchers can apply for up to $10,000 in free research credits.
Explore Purna's Molecular Intelligence Platform
AI-powered workspace for biology teams to accelerate drug discovery from target identification to lead optimization.
Try Purna AI →
